The ggRandomForests package extracts tidy data objects from either randomForestSRC or randomForest fits and feeds them into familiar ggplot2 workflows. This vignette highlights the most common objects— gg_error, gg_variable, and gg_vimp—along with a small helper for building balanced conditioning intervals.
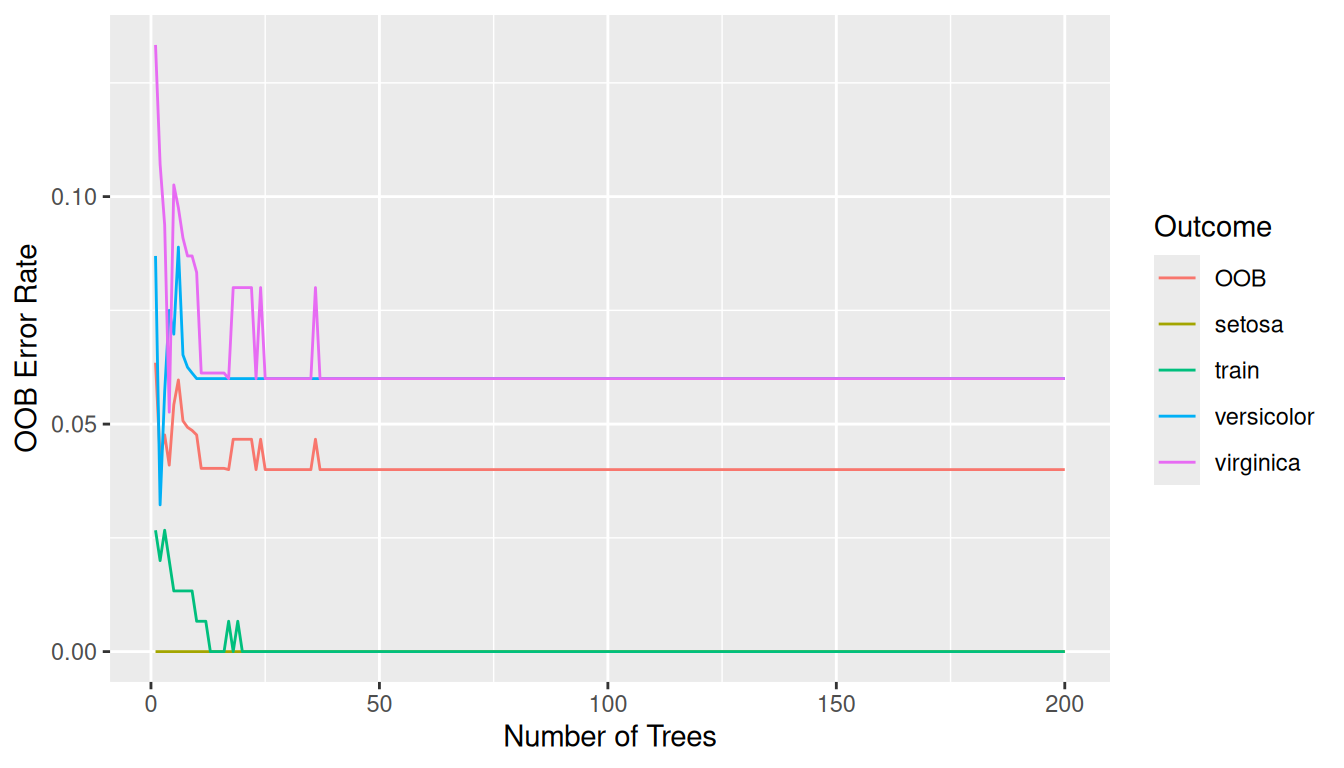
Error trajectories with gg_error()
OOB setosa versicolor virginica ntree train
1 0.06349206 0 0.08695652 0.13333333 1 0.02666667
2 0.04255319 0 0.03225806 0.10714286 2 0.02000000
3 0.04761905 0 0.05714286 0.09375000 3 0.02666667
4 0.04098361 0 0.07500000 0.05263158 4 0.02000000
5 0.05426357 0 0.06976744 0.10256410 5 0.01333333
6 0.05970149 0 0.08888889 0.09756098 6 0.01333333
The gg_error() object stores the cumulative OOB error rate for each outcome column plus the ntree counter. When training = TRUE, the function reconstructs the original model frame and appends the in-bag error trajectory (train). Plotting overlays both curves by default:
Marginal dependence via gg_variable()
Classes 'gg_variable', 'regression' and 'data.frame': 506 obs. of 2 variables:
$ lstat: num 4.98 9.14 4.03 2.94 5.33 ...
$ yhat : num 29.2 22.5 35.1 36.4 33.4 ...
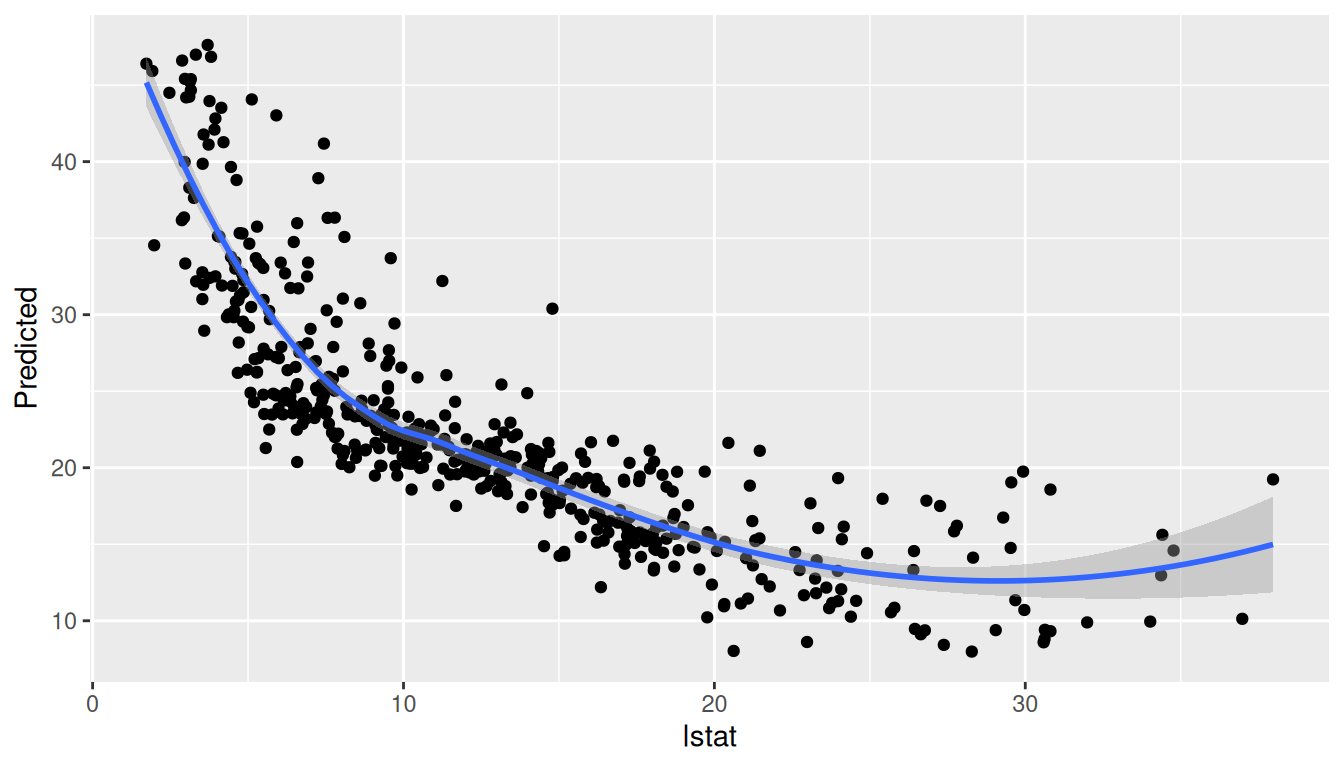
Because the original training data are recovered from the model call, gg_variable() works even when the forest was trained within helper functions or against a subset() expression. The output keeps the raw predictors plus either a continuous yhat column (regression) or per-class probabilities (yhat.<class> for classification). Plotting a single variable is straightforward:
plot(var_df, xvar = "lstat")
`geom_smooth()` using method = 'loess' and formula = 'y ~ x'
Survival forests can request multiple horizons using the time argument; non-OOB predictions are available by setting oob = FALSE.
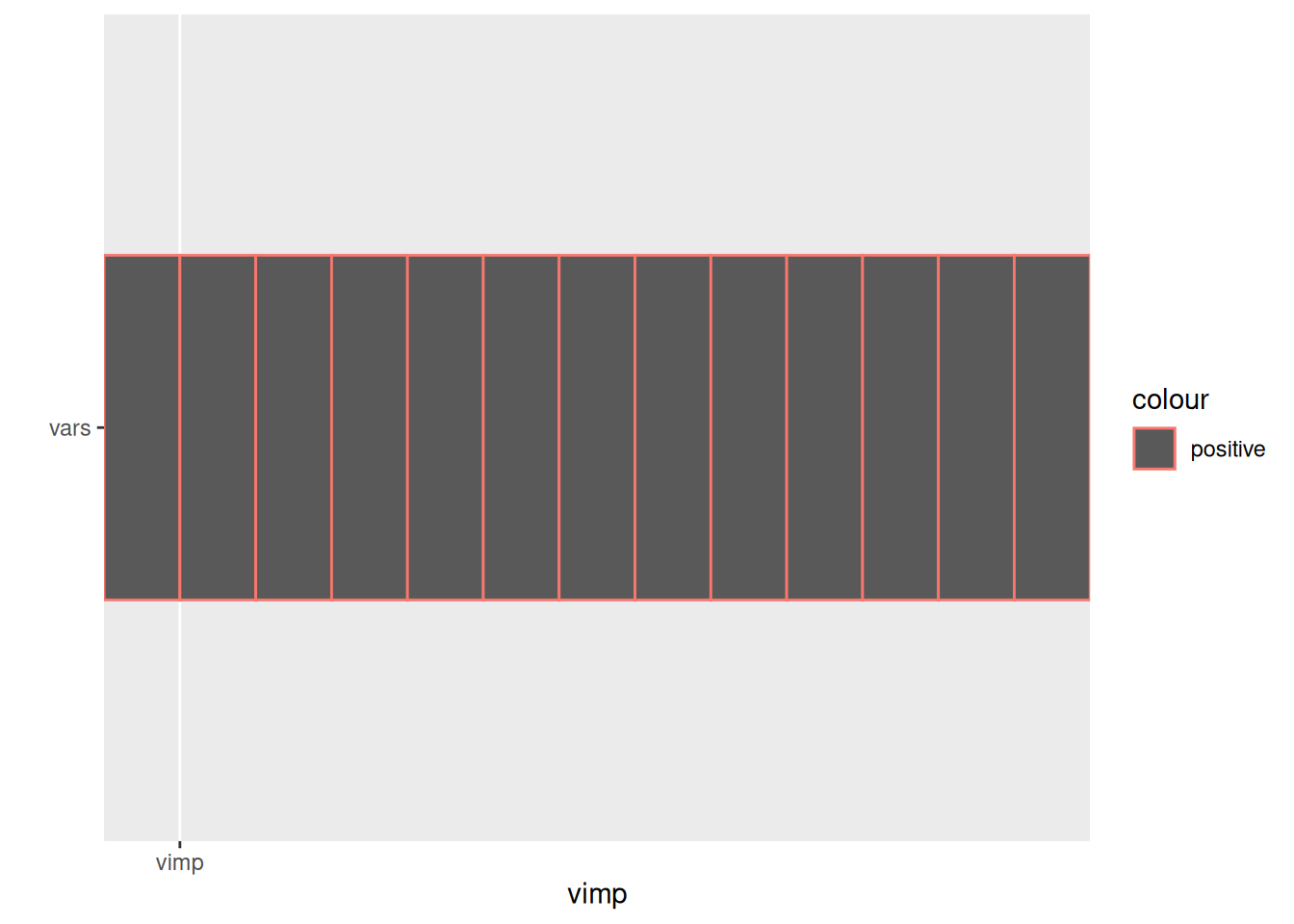
Variable importance with gg_vimp()
vimp_df <- ggRandomForests::gg_vimp(rf_boston)
head(vimp_df)
# A tibble: 6 × 4
vars set vimp positive
<fct> <chr> <dbl> <lgl>
1 lstat vimp 13005. TRUE
2 rm vimp 11662. TRUE
3 dis vimp 2849. TRUE
4 indus vimp 2751. TRUE
5 ptratio vimp 2698. TRUE
6 crim vimp 2646. TRUE
If a randomForest object lacks stored importance scores, gg_vimp() tries to compute them on the fly. When the forest truly cannot provide the information (for example when importance = FALSE and the predictors are no longer accessible), the function emits a warning and returns NA placeholders so plots still render.
Balanced conditioning cuts with quantile_pts()
rm_breaks <- ggRandomForests::quantile_pts(boston$rm, groups = 6, intervals = TRUE)
rm_groups <- cut(boston$rm, breaks = rm_breaks)
table(rm_groups)
rm_groups
(3.56,5.76] (5.76,5.99] (5.99,6.21] (6.21,6.44] (6.44,6.85] (6.85,8.78]
85 84 84 85 84 84
The helper wraps stats::quantile() to produce evenly populated strata that drop directly into cut() when building coplots or facet labels.