Validates a wide cluster-assignment data frame, resolves node level
ordering, computes default node colours if not supplied, and pre-computes
the long-format Sankey data. Call plot.hv_sankey on the
result to obtain a bare ggplot2 Sankey diagram using
ggsankey geoms.
Usage
hv_sankey(
data,
cluster_cols = paste0("C", 2:9),
node_levels = NULL,
node_colours = NULL
)Arguments
- data
Data frame; one row per patient. Must contain all columns named in
cluster_cols.- cluster_cols
Character vector of column names giving the cluster assignments at each value of K, in ascending K order. Default
paste0("C", 2:9).- node_levels
Character vector giving the display order of node labels (bottom to top within each column). If
NULL(default), the existing factor levels of the first cluster column are used.- node_colours
Named character vector mapping node labels to fill colours. If
NULL(default), colours are drawn fromRColorBrewer::brewer.pal(9, "Set1")in the orderc(2, 6, 8, 4, 3, 5, 7, 1, 9).
Value
An object of class c("hv_sankey", "hv_data"):
$dataThe long-format Sankey data frame (four columns:
x,node,next_x,next_node).$metaNamed list:
cluster_cols,node_levels,node_colours,n_patients,n_k.$tablesEmpty list.
Examples
dta <- sample_cluster_sankey_data(n = 300, seed = 42)
if (requireNamespace("ggsankey", quietly = TRUE)) {
# 1. Build data object
sn <- hv_sankey(dta)
sn # prints cluster cols and node count
# 2. Bare plot -- undecorated ggplot returned by plot.hv_sankey
p <- plot(sn)
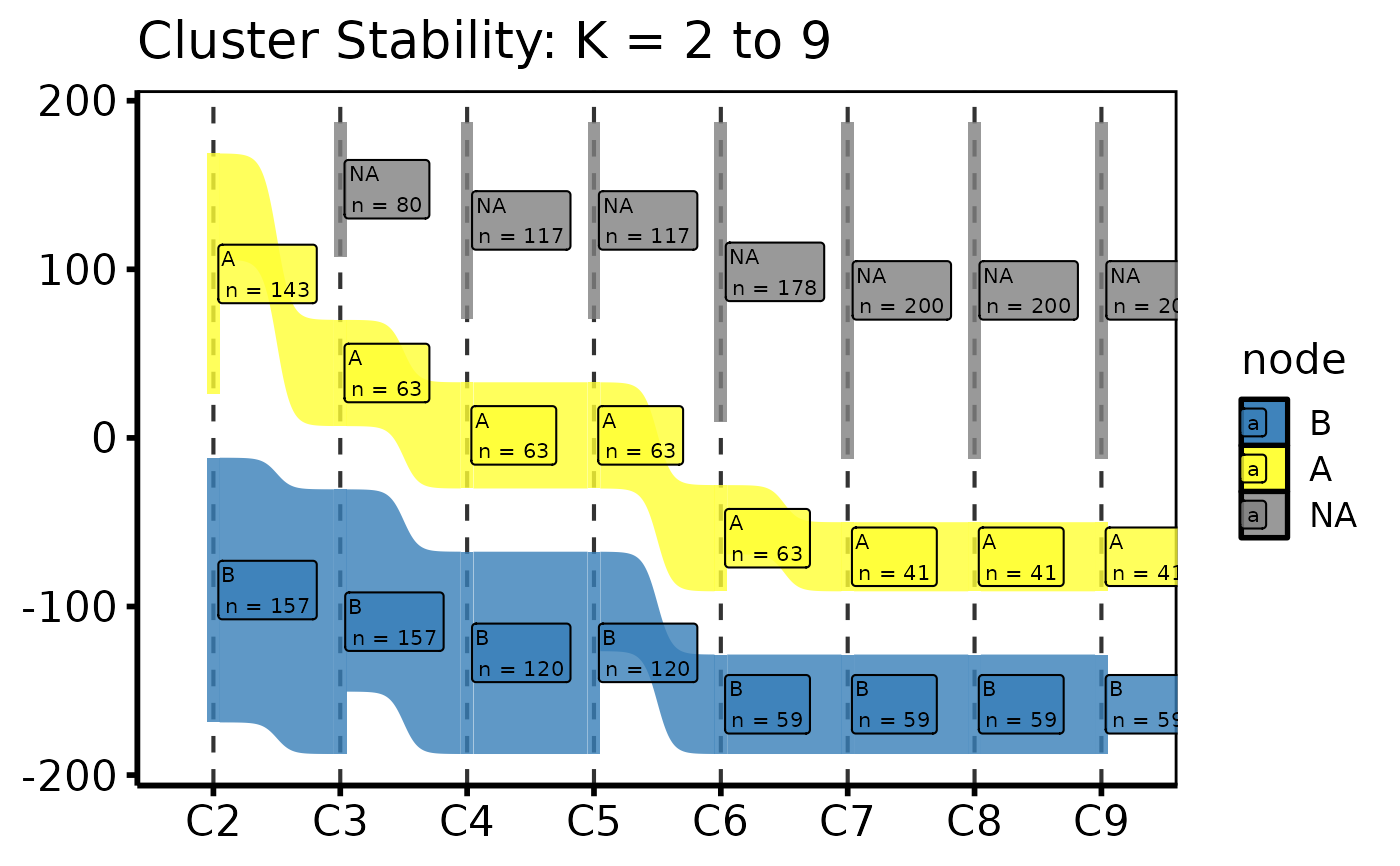
# 3. Decorate: axis labels and theme
p +
ggplot2::labs(x = NULL, title = "Cluster Stability: K = 2 to 9") +
hv_theme("poster")
}
#> Warning: The `size` argument of `element_rect()` is deprecated as of ggplot2 3.4.0.
#> ℹ Please use the `linewidth` argument instead.
#> ℹ The deprecated feature was likely used in the ggsankey package.
#> Please report the issue at <https://github.com/davidsjoberg/ggsankey/issues>.