Overview
This guide maps every SAS template in the HVTI statistics group template library to its R equivalent in the hvtiPlotR package. If you know the SAS template name (e.g., tp.np.afib.ivwristm.avrg_curv.binary.sas), look it up in the table below and jump to the corresponding section for a working R example.
The guide is organized by template family (the two-letter prefix after tp.). New ports are added to this document as they become available.
Key concepts for SAS users
ggplot2 builds plots in layers. Instead of one macro call with many color=, xaxis=, and footnote= options, you chain + operations:
hv_theme() replaces SAS device= / style options. Use "dark_ppt" / "ppt" for PowerPoint slides, "light_ppt" for light-background slides, "manuscript" for journal figures, and "poster" for conference posters.
Functions return a ggplot object; they do not display it. Call the plot object at the top level (or use print()) to render it, or save with ggsave() / save_ppt().
Pre-fitted models, not raw data. Most functions accept the output of a statistical model (curve datasets, probability estimates) rather than individual patient records — mirroring the SAS workflow where %decompos() or %kaplan computes estimates and a separate template step produces the figure.
Nonparametric temporal trends (tp.np.*)
All tp.np.* templates share a common SAS workflow:
- Fit patient-specific temporal profiles with
%decompos().
- Average profiles across patients with
PROC SUMMARY → mean_curv dataset.
- Optionally compute bootstrap confidence intervals →
boots_ci dataset.
- Compute binned patient-level summaries (deciles/quintiles) →
means dataset.
- Call the plotting template.
The R port replaces step 5. Steps 1–4 still run in SAS; export the resulting datasets to CSV and read them into R:
curve_data <- read.csv("mean_curv.csv") # or boots_ci if CI bands are needed
data_pts <- read.csv("means.csv") # optional binned data points
SAS column name mapping
iv_echo / iv_wristm / iv_fyrs
|
x_col |
Time variable |
prev / mnprev / _p_ / est
|
estimate_col |
Curve estimate |
cll_p68 / cll_p95
|
lower_col |
CI lower band |
clu_p68 / clu_p95
|
upper_col |
CI upper band |
Grouping variable (e.g., group) |
group_col |
NULL for single curve |
mtime / mmtime (means dataset) |
dp_x_col |
Binned point x |
mprev / mnprev (means dataset) |
dp_y_col |
Binned point y |
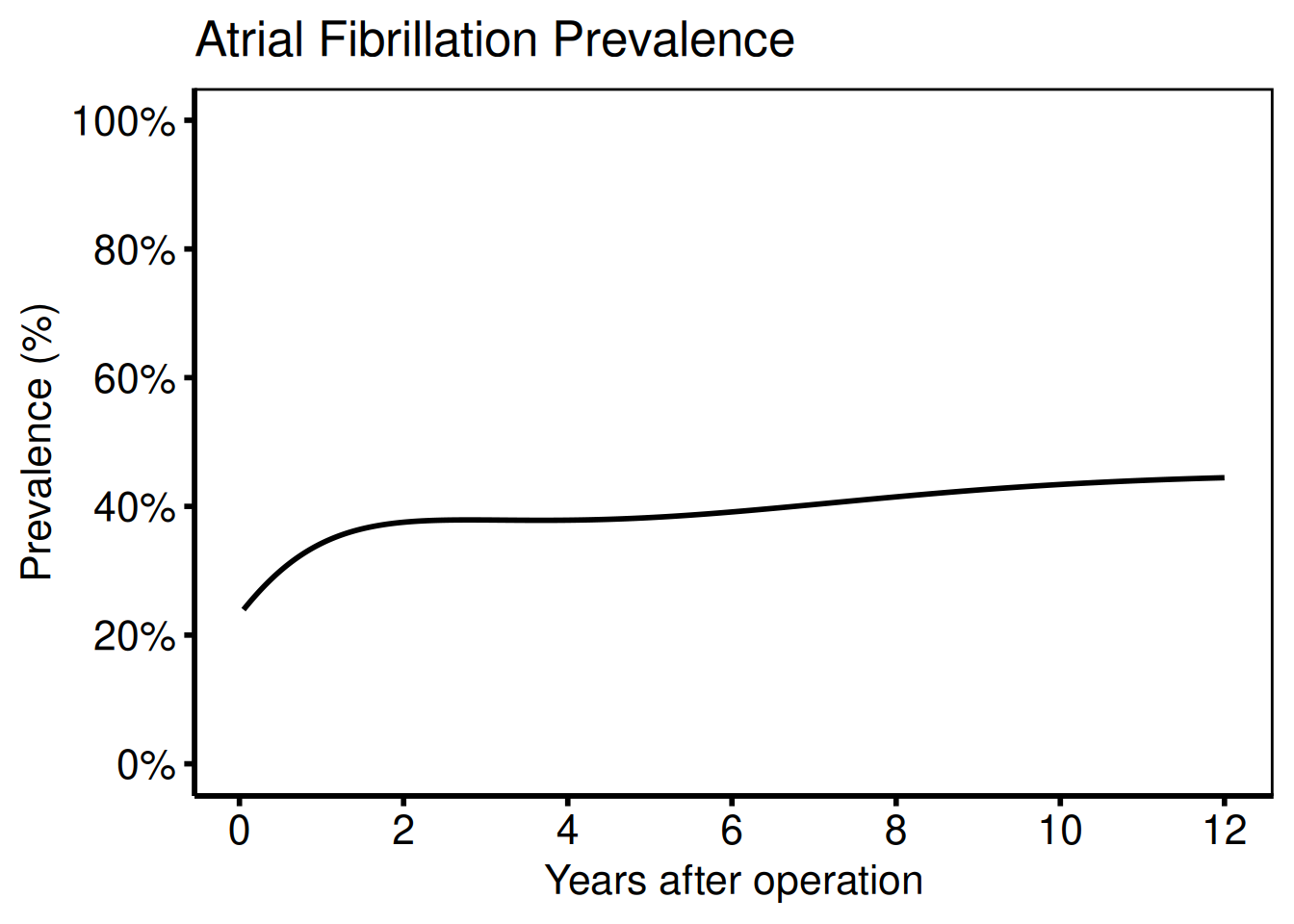
Average curve — binary outcome
Ports: tp.np.afib.ivwristm.avrg_curv.binary.sas, tp.np.afib.ivwristm.pt_specific.binary.sas, tp.np.tr.icdpr.avg_curv.sas
dat <- sample_nonparametric_curve_data(
n = 500,
time_max = 12,
outcome_type = "probability"
)
dat_pts <- sample_nonparametric_curve_points(n = 500, time_max = 12)
# Minimal plot
plot(hv_nonparametric(dat)) +
scale_x_continuous(breaks = seq(0, 12, 2)) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Prevalence (%)",
title = "Atrial Fibrillation Prevalence"
) +
hv_theme("poster")
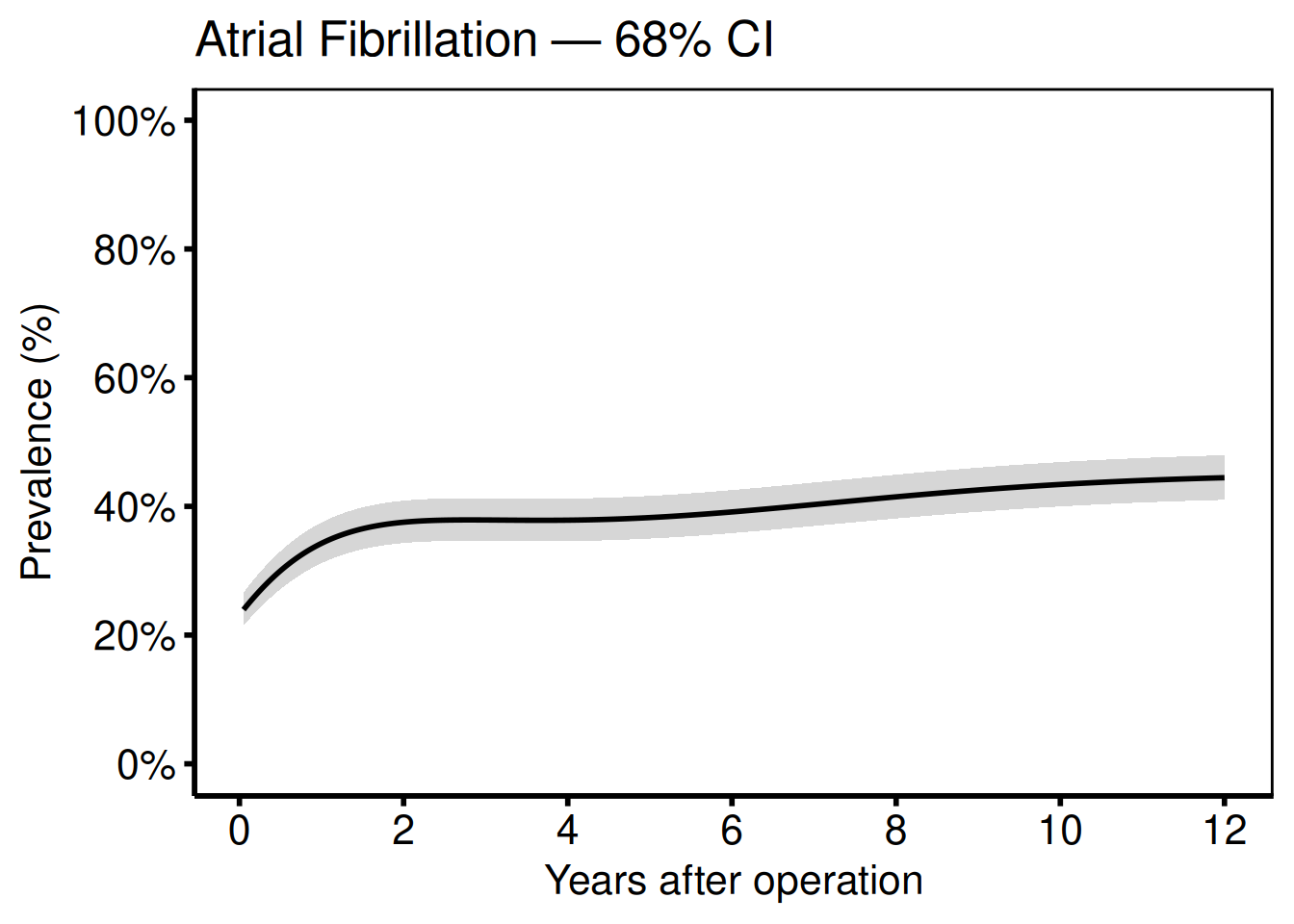
Add 68% CI bands (one standard error; matches SAS boots_ci with cll_p68 / clu_p68):
plot(hv_nonparametric(
dat,
lower_col = "lower",
upper_col = "upper"
)) +
scale_x_continuous(breaks = seq(0, 12, 2)) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Prevalence (%)",
title = "Atrial Fibrillation — 68% CI"
) +
hv_theme("poster")
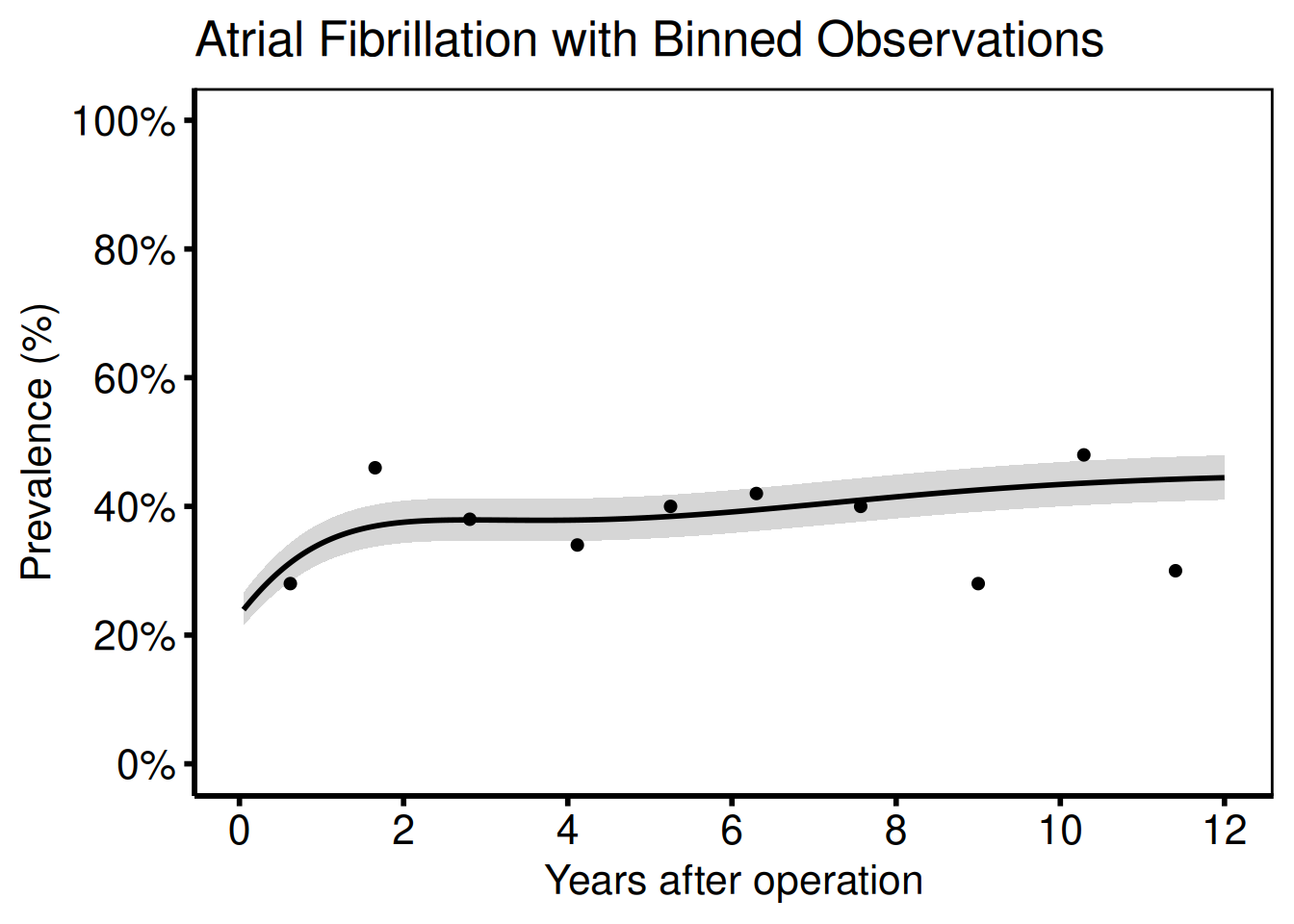
Add binned data summary points (matches the SAS means dataset):
plot(hv_nonparametric(
dat,
lower_col = "lower",
upper_col = "upper",
data_points = dat_pts
)) +
scale_x_continuous(breaks = seq(0, 12, 2)) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Prevalence (%)",
title = "Atrial Fibrillation with Binned Observations"
) +
hv_theme("poster")
Continuous outcome
Ports: tp.np.fev.double.univariate.continuous.sas, tp.np.fev.u.trend.continuous.sas
For continuous outcomes (FEV1, AV peak gradient) set outcome_type = "continuous". The y-axis label and scale change accordingly; everything else is identical.
dat_cont <- sample_nonparametric_curve_data(
n = 400,
time_max = 10,
outcome_type = "continuous"
)
dat_cont_pts <- sample_nonparametric_curve_points(
n = 400,
time_max = 10,
outcome_type = "continuous"
)
plot(hv_nonparametric(
dat_cont,
lower_col = "lower",
upper_col = "upper",
data_points = dat_cont_pts
)) +
scale_x_continuous(breaks = seq(0, 10, 2)) +
scale_y_continuous(limits = c(0, 4)) +
labs(
x = "Years after operation",
y = expression(FEV[1] ~ (L)),
title = "FEV\u2081 Temporal Trend"
) +
annotate("text",
x = 9, y = 3.5,
label = "68% CI band",
hjust = 1, size = 3
) +
hv_theme("poster")
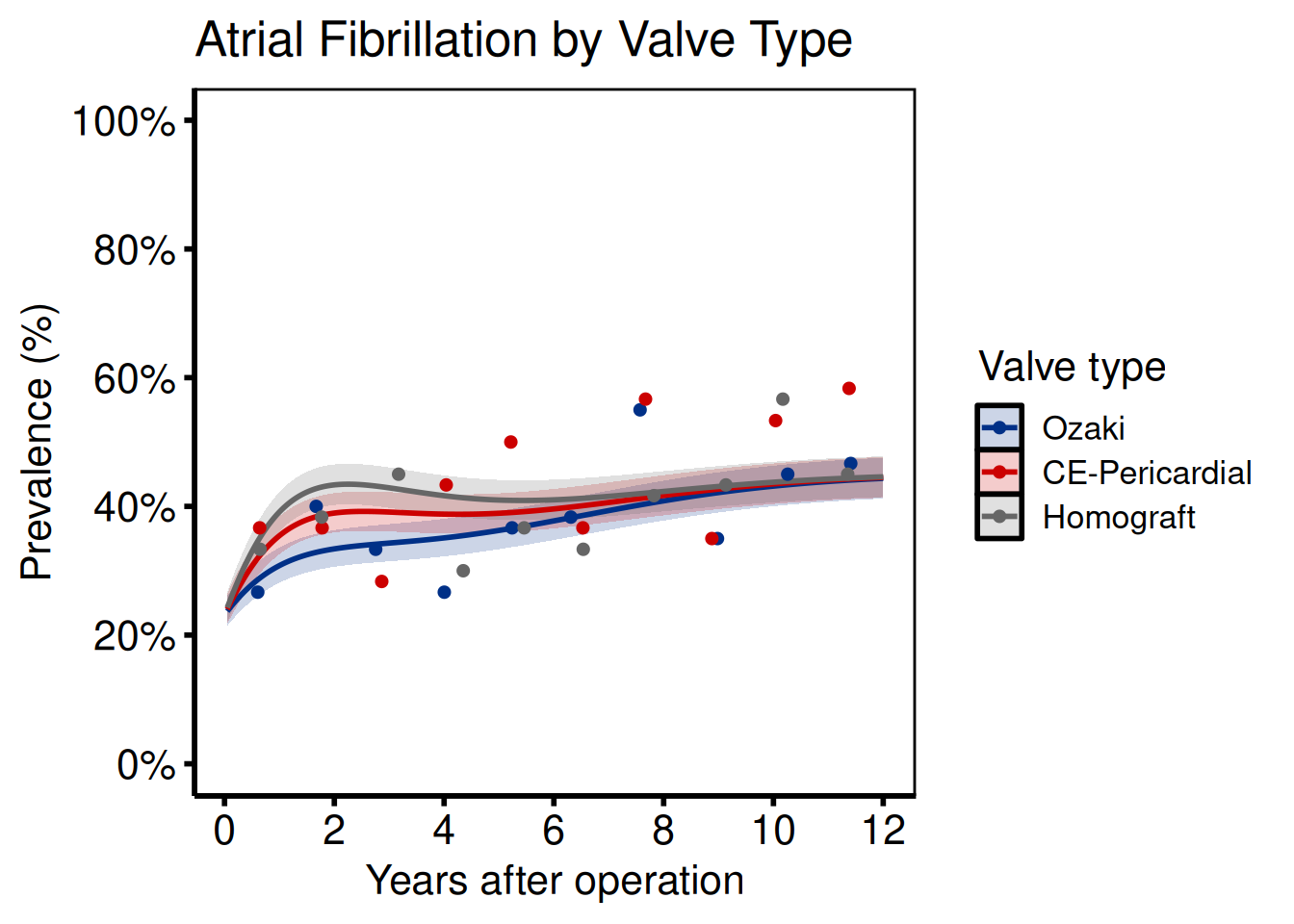
Multi-group comparison
Ports: tp.np.afib.mult.avrg_curv.binary.sas, tp.np.afib.mult.pt_spec.binary.sas, tp.np.avpkgrad_ozak_ind_mtwt.sas, tp.np.fev.multivariate.continuous.sas
Provide a named vector to groups when generating sample data, or set group_col when supplying your own curve data.
grp_def <- c("Ozaki" = 0.7, "CE-Pericardial" = 1.1, "Homograft" = 1.4)
dat_grp <- sample_nonparametric_curve_data(
n = 600,
time_max = 12,
groups = grp_def
)
dat_grp_pts <- sample_nonparametric_curve_points(
n = 600,
time_max = 12,
groups = grp_def
)
plot(hv_nonparametric(
dat_grp,
group_col = "group",
lower_col = "lower",
upper_col = "upper",
data_points = dat_grp_pts
)) +
scale_colour_manual(
values = c("Ozaki" = "#003087", "CE-Pericardial" = "#CC0000", "Homograft" = "#666666")
) +
scale_fill_manual(
values = c("Ozaki" = "#003087", "CE-Pericardial" = "#CC0000", "Homograft" = "#666666")
) +
scale_x_continuous(breaks = seq(0, 12, 2)) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Prevalence (%)",
colour = "Valve type",
fill = "Valve type",
title = "Atrial Fibrillation by Valve Type"
) +
hv_theme("poster")
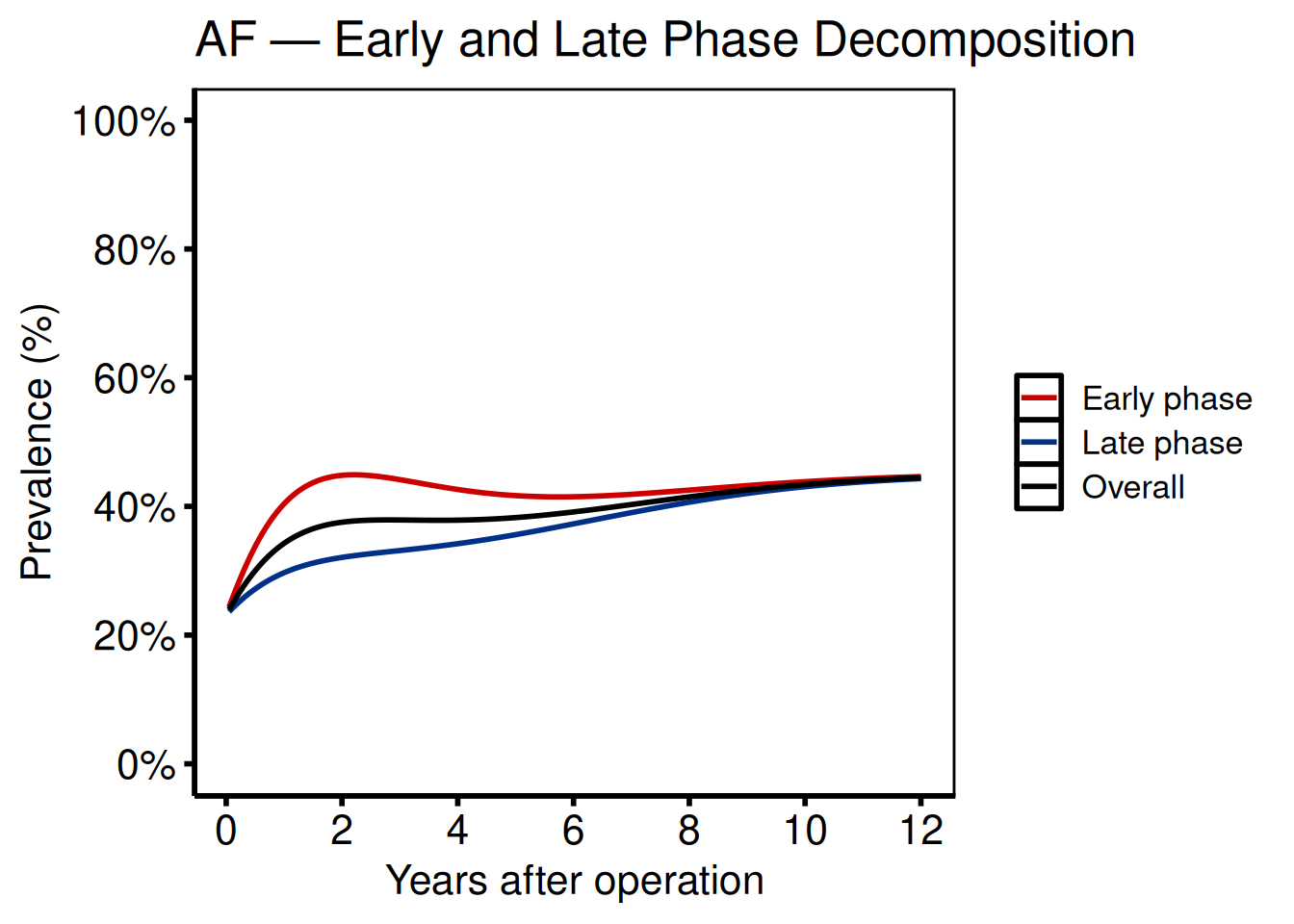
Phase decomposition
Ports: tp.np.afib.ivwristm.pt_spec_phases.binary.sas, tp.np.tr.ivecho.u.phases.sas
Phase plots separate the early (bell-shaped, incomplete hazard) and late (Weibull CDF, complete hazard) components. Supply a group_col whose levels label the phases.
dat_phase <- sample_nonparametric_curve_data(
n = 500,
groups = c("Early phase" = 1.5, "Late phase" = 0.6, "Overall" = 1.0)
)
plot(hv_nonparametric(
dat_phase,
group_col = "group"
)) +
scale_colour_manual(
values = c("Early phase" = "#CC0000",
"Late phase" = "#003087",
"Overall" = "black")
) +
scale_linetype_manual(
values = c("Early phase" = "dashed",
"Late phase" = "dashed",
"Overall" = "solid")
) +
scale_x_continuous(breaks = seq(0, 12, 2)) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Prevalence (%)",
colour = NULL,
title = "AF — Early and Late Phase Decomposition"
) +
hv_theme("poster")
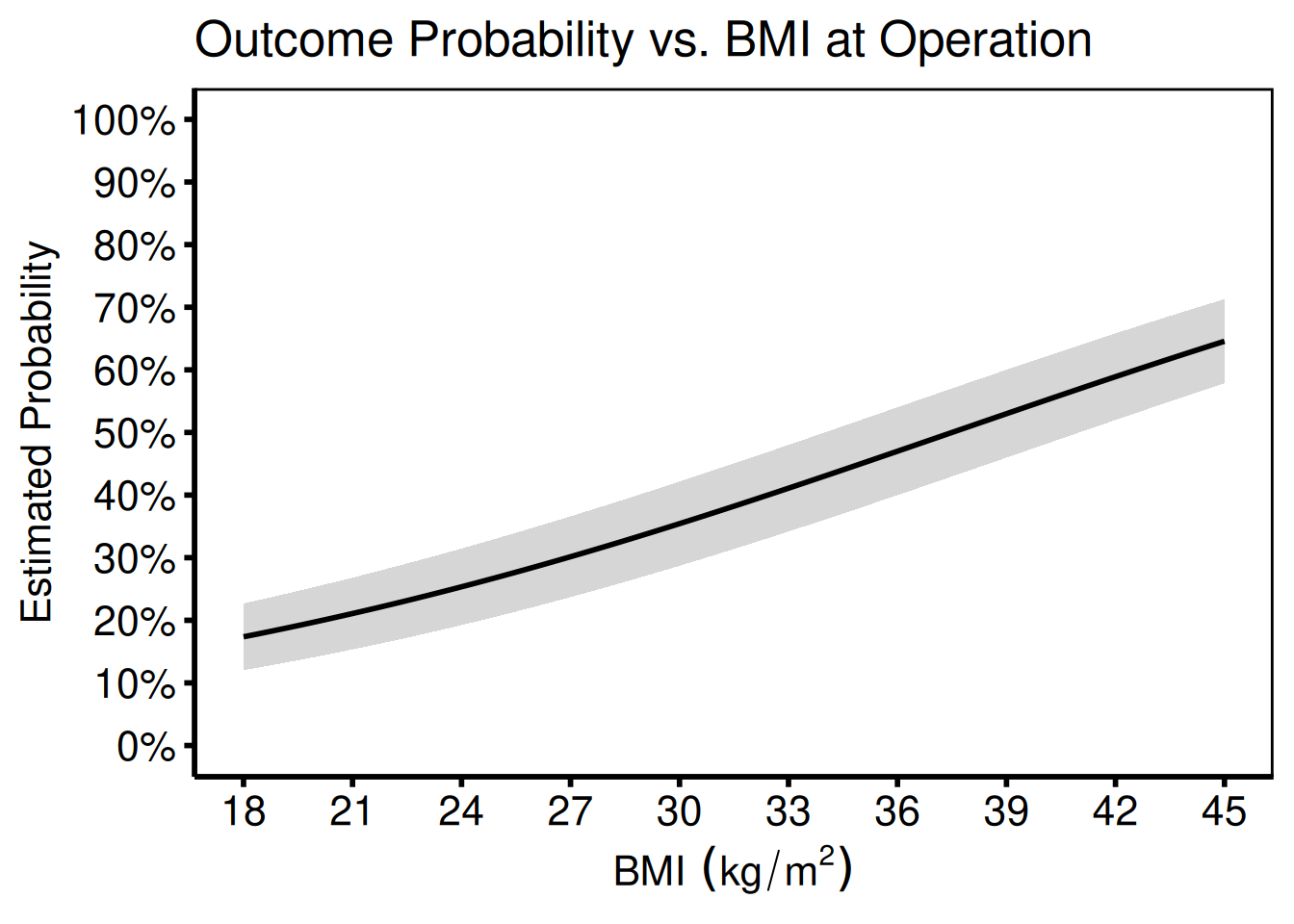
Covariate on the x-axis
Port: tp.np.z0axdpo.continuous.bmi_xaxis.sas
When BMI or another continuous covariate (rather than time) is on the x-axis, the function signature is identical — simply pass the covariate column name to x_col.
# Simulate covariate (BMI) x-axis data
set.seed(42)
n_pts <- 300
bmi <- seq(18, 45, length.out = n_pts)
est <- plogis(-3 + 0.08 * bmi)
se <- sqrt(est * (1 - est) / 50)
bmi_curve <- data.frame(
bmi = bmi,
est = est,
lower = pmax(0, est - qnorm(0.84) * se),
upper = pmin(1, est + qnorm(0.84) * se)
)
plot(hv_nonparametric(
bmi_curve,
x_col = "bmi",
estimate_col = "est",
lower_col = "lower",
upper_col = "upper"
)) +
scale_x_continuous(
breaks = seq(18, 45, 3),
limits = c(18, 45)
) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.1),
labels = scales::percent
) +
labs(
x = expression(BMI ~ (kg/m^2)),
y = "Estimated Probability",
title = "Outcome Probability vs. BMI at Operation"
) +
hv_theme("poster")
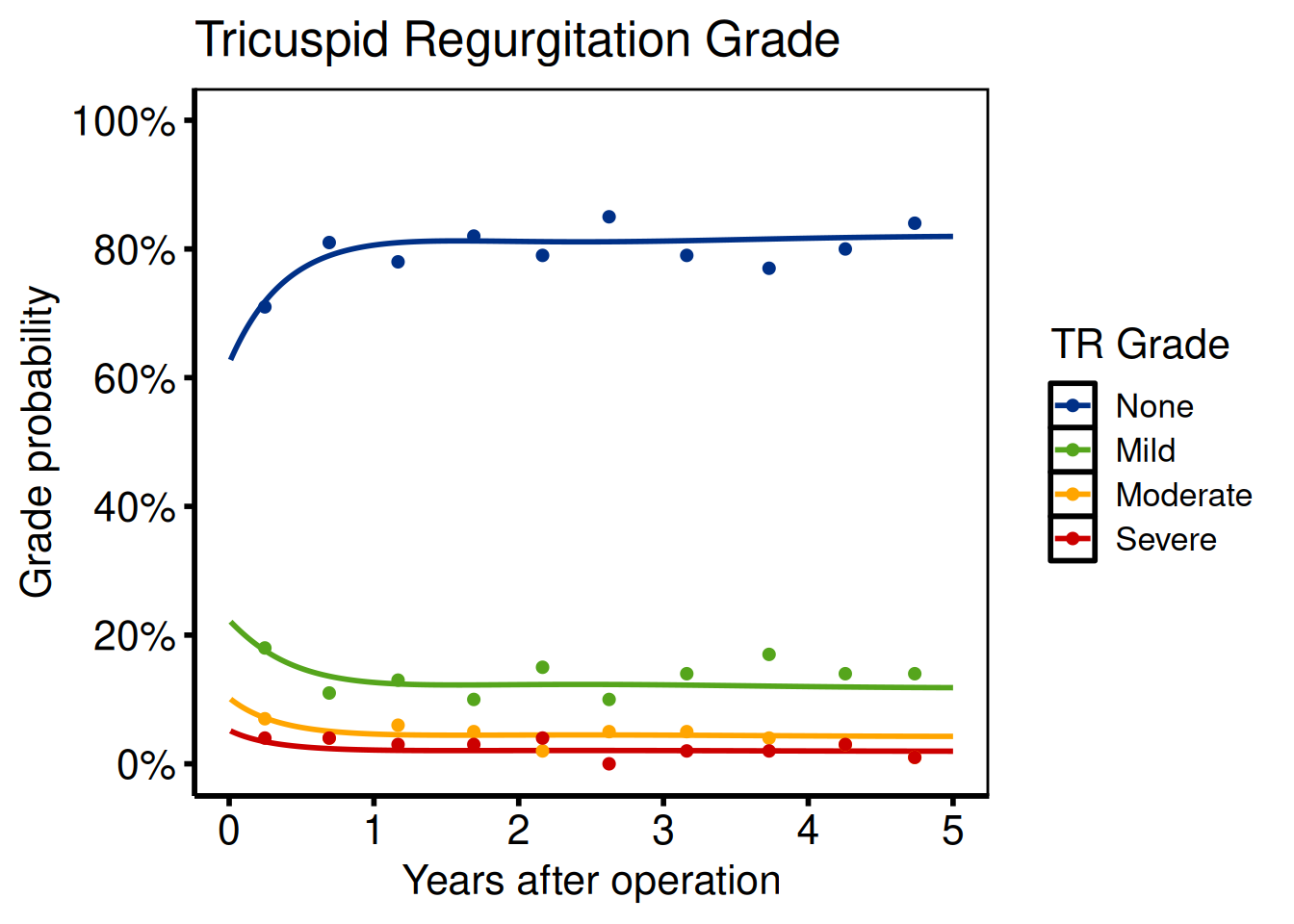
Ordinal outcomes
Port: tp.np.tr.ivecho.average_curv.ordinal.sas
Ordinal templates (TR grade, AR grade) use hv_ordinal(). The critical migration step is reshaping the SAS wide-format predict dataset (one column per grade) to long format.
SAS reshape (do this before reading into R):
/* SAS wide format: p0, p1, p2, p3 */
data predict_wide;
set predict;
run;
R reshape:
ord_labels <- c("None", "Mild", "Moderate", "Severe")
dat_ord <- sample_nonparametric_ordinal_data(
n = 1000,
time_max = 5,
grade_labels = ord_labels
)
dat_ord_pts <- sample_nonparametric_ordinal_points(
n = 1000,
time_max = 5,
grade_labels = ord_labels
)
plot(hv_ordinal(
dat_ord,
grade_col = "grade",
data_points = dat_ord_pts
)) +
scale_colour_manual(
values = c(
"None" = "#003087",
"Mild" = "#55A51C",
"Moderate" = "#FFA500",
"Severe" = "#CC0000"
)
) +
scale_x_continuous(breaks = 0:5) +
scale_y_continuous(
limits = c(0, 1),
breaks = seq(0, 1, 0.2),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Grade probability",
colour = "TR Grade",
title = "Tricuspid Regurgitation Grade"
) +
hv_theme("poster")
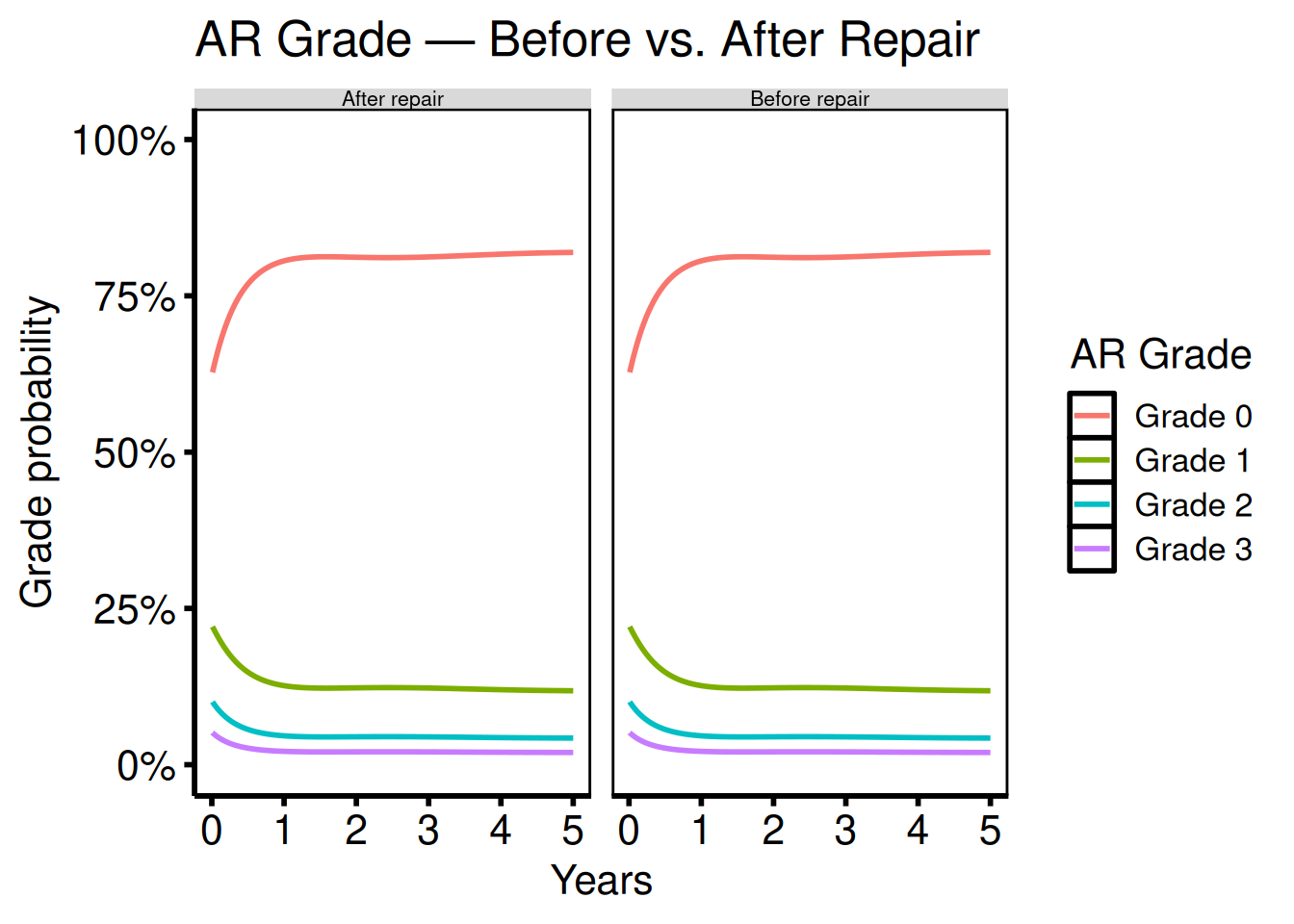
Ordinal multi-scenario comparison
Port: tp.np.po_ar.u_multi.ordinal.sas
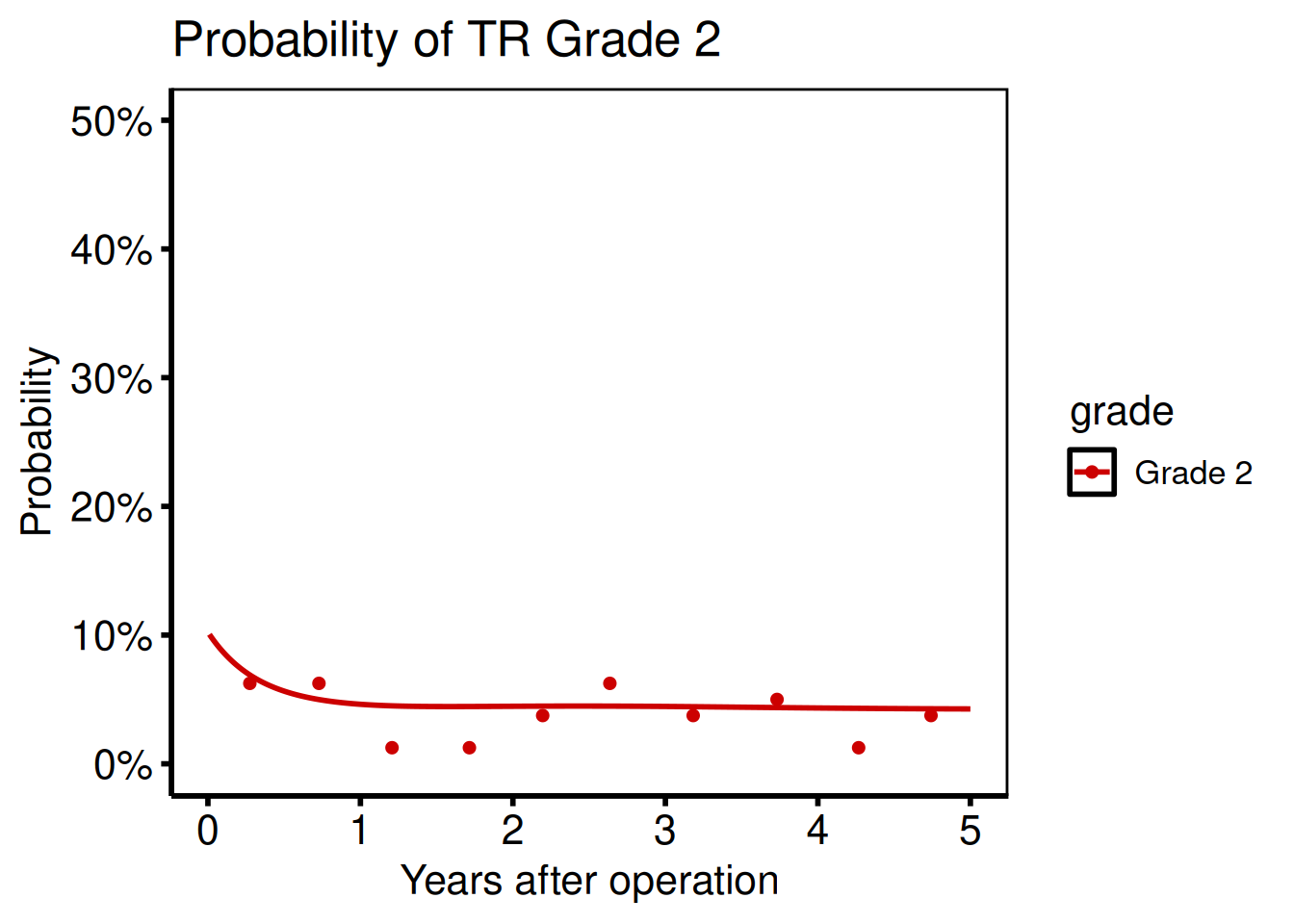
Ordinal phase independence
Port: tp.np.tr.ivecho.independence.sas
To examine a single grade in isolation, filter the long-format curve before passing it to the function:
dat_ind <- sample_nonparametric_ordinal_data(n = 800)
pts_ind <- sample_nonparametric_ordinal_points(n = 800)
grade_2 <- dat_ind[dat_ind$grade == "Grade 2", ]
dp_grade_2 <- pts_ind[pts_ind$grade == "Grade 2", ]
plot(hv_ordinal(
grade_2,
grade_col = "grade",
data_points = dp_grade_2
)) +
scale_colour_manual(values = c("Grade 2" = "#CC0000")) +
scale_x_continuous(breaks = seq(0, 5, 1)) +
scale_y_continuous(
limits = c(0, 0.5),
labels = scales::percent
) +
labs(
x = "Years after operation",
y = "Probability",
title = "Probability of TR Grade 2"
) +
hv_theme("poster")
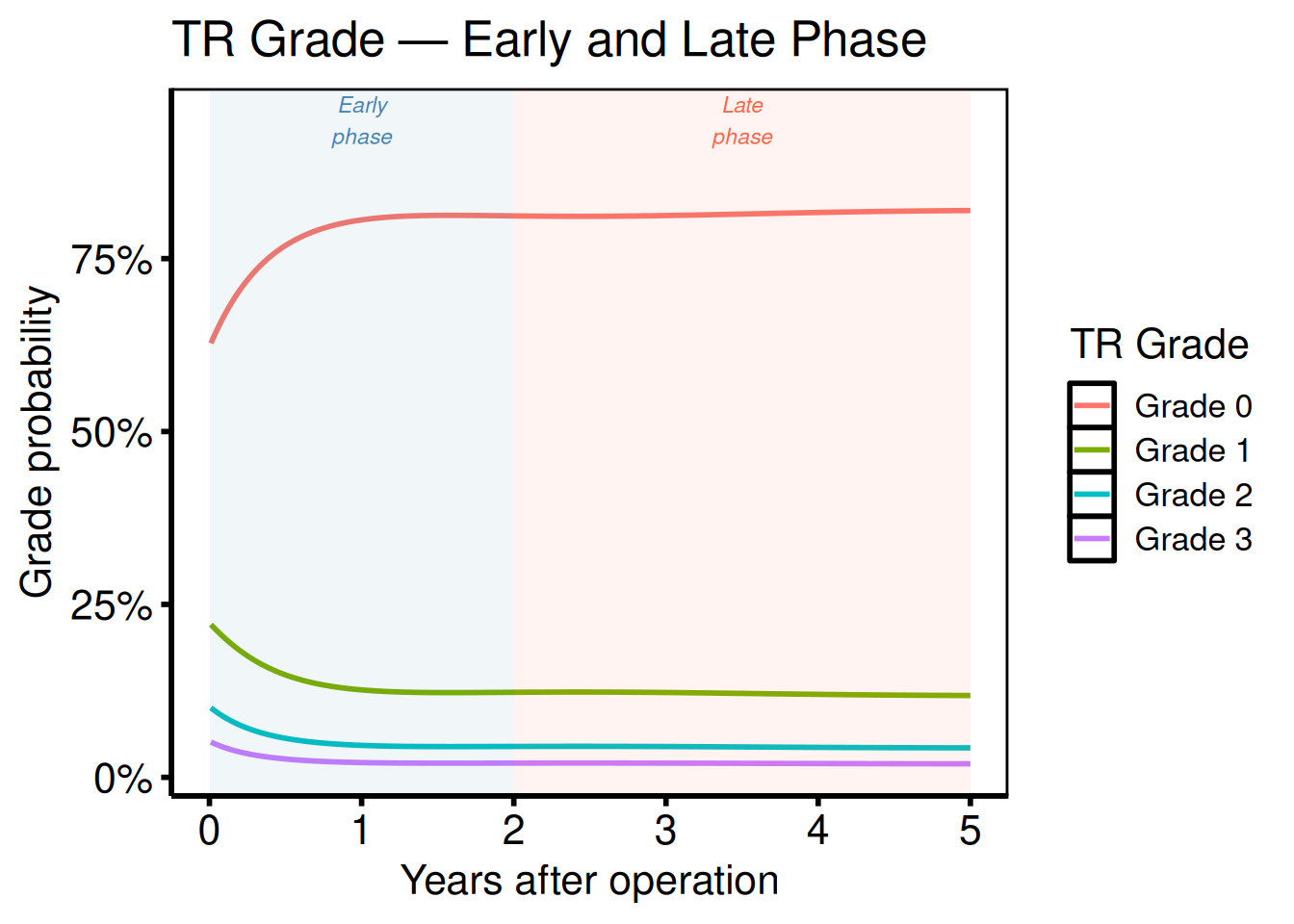
Ordinal phases
Port: tp.np.tr.ivecho.u.phases.sas
Phase-decomposed ordinal plots combine phase labels with grade colours. Create the figure by plotting grade-specific curves and annotating early vs. late phase regions:
dat_ph <- sample_nonparametric_ordinal_data(n = 800, seed = 7)
plot(hv_ordinal(dat_ph, grade_col = "grade")) +
annotate("rect",
xmin = 0, xmax = 2, ymin = -Inf, ymax = Inf,
fill = "steelblue", alpha = 0.07
) +
annotate("text",
x = 1, y = 0.95, label = "Early\nphase",
size = 3, colour = "steelblue", fontface = "italic"
) +
annotate("rect",
xmin = 2, xmax = 5, ymin = -Inf, ymax = Inf,
fill = "tomato", alpha = 0.07
) +
annotate("text",
x = 3.5, y = 0.95, label = "Late\nphase",
size = 3, colour = "tomato", fontface = "italic"
) +
scale_x_continuous(breaks = 0:5) +
scale_y_continuous(labels = scales::percent) +
labs(
x = "Years after operation",
y = "Grade probability",
colour = "TR Grade",
title = "TR Grade — Early and Late Phase"
) +
hv_theme("poster")
Survival analysis (tp.ac.dead.*, tp.cp.dead.*)
Ports: tp.ac.dead.sas, tp.cp.dead.sas
R equivalent: hv_survival()
hv_survival() wraps survfit() from the survival package and returns an S3 object. Call plot(km, type = ...) to render one of the five plot types matching the SAS %kaplan / %nelsont macro output flags (PLOTS, PLOTC, PLOTH, PLOTL). Tidy data frames are accessible via km$tables.
dta <- sample_survival_data(n = 500, seed = 42)
km <- hv_survival(dta)
# Kaplan–Meier survival (PLOTS)
plot(km) +
scale_y_continuous(
breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")
) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(
x = "Years after operation",
y = "Survival (%)",
title = "Freedom from Death"
) +
hv_theme("poster")
# Cumulative hazard (PLOTC)
plot(km, type = "cumhaz") +
labs(x = "Years", y = "Cumulative Hazard") +
hv_theme("poster")
# Hazard rate (PLOTH)
plot(km, type = "hazard") +
labs(x = "Years", y = "Instantaneous Hazard") +
hv_theme("poster")
# Log-log survival (PH check)
plot(km, type = "loglog") +
labs(x = "log(Years)", y = "log(-log S(t))") +
hv_theme("poster")
# Integrated survivorship (PLOTL)
plot(km, type = "life") +
labs(x = "Years", y = "Restricted Mean Survival (years)") +
hv_theme("poster")
# Access the KM data, risk table, and report table
km$data # tidy KM data frame
km$tables$risk # numbers-at-risk table
km$tables$report # survival at report times
SAS → R argument mapping
data= |
data |
Patient-level data frame |
time= |
time_col |
Time-to-event column name |
event= |
event_col |
Event indicator column (1 = event) |
group= |
group_col |
Stratification variable |
method=kaplan |
method = "kaplan-meier" |
Default |
method=nelsont |
method = "nelson-aalen" |
Fleming–Harrington |
alpha=0.05 |
conf_level = 0.95 |
Default is 0.95 |
tp=0 1 2 3 5 7 10 |
report_times |
Table time points |
Goodness of follow-up (tp.dp.gfup.R)
Port: tp.dp.gfup.R
R equivalent: hv_followup()
Covariate balance (tp.lp.propen.cov_balance.R)
Port: tp.lp.propen.cov_balance.R
R equivalent: hv_balance()
The SAS export arrives wide (one column per time-point). Reshape to long format before calling hv_balance(). Pass var_levels to control the bottom-to-top display order of covariates (matches ylabel in the original script).
# Reshape wide → long (mirrors the original script)
ylabel <- c(
"Cardiac Output", "BAV", "Cardiac Index", "NYHA -Functional Class",
"COPD-Oxy", "Date of Surg", "Creatinine", "Age", "CAD",
"Female", "LV PWT", "Mismatch", "HTN", "PVD", "Sinitibular Jcn: diam"
)
names(dta) <- c("variable", "Before match", "After match")
dta_long <- reshape(
dta,
direction = "long",
varying = c("Before match", "After match"),
v.names = "std_diff",
timevar = "group",
times = c("Before match", "After match"),
idvar = "variable"
)
n_vars <- length(ylabel)
cb <- hv_balance(
data = dta_long,
variable_col = "variable",
group_col = "group",
std_diff_col = "std_diff",
var_levels = ylabel
)
stdifPlot <- plot(cb) +
scale_color_manual(
values = c("Before match" = "red4", "After match" = "blue3"),
name = NULL
) +
scale_shape_manual(
values = c("Before match" = 17L, "After match" = 15L),
name = NULL
) +
scale_x_continuous(limits = c(-40, 30), breaks = seq(-40, 30, 10)) +
labs(x = "Standardized difference: SAVR - TF:TAVR (%)", y = "") +
annotate("text", x = -30, y = 0, label = "More likely TF-TAVR", size = 4.5) +
annotate("text", x = 22, y = n_vars, label = "More likely SAVR", size = 4.5) +
theme(legend.position = c(0.20, 0.935)) +
hv_theme("poster")
ggsave(here::here("graphs", "lp_cov-balance-SAVR_TF-TAVR.pdf"),
plot = stdifPlot, height = 7, width = 8)
Alluvial flow (tp.dp.female_bicus_preAR_sankey.R)
Port: tp.dp.female_bicus_preAR_sankey.R
R equivalent: hv_alluvial()
Cluster stability Sankey (PAM analysis)
Source: PAM clustering analysis pipeline (R script using ggsankey)
R equivalent: hv_sankey()
hv_sankey() draws a Sankey diagram showing how patients flow between letter-labelled clusters as the number of clusters K increases from 2 to 9. Each column is one K; flow bands show assignment changes between consecutive K values; node labels show cluster letter + patient count.
The original code used ggsankey::make_long() with hard-coded column names and ordering vectors. The R port accepts any cluster columns and ordering via cluster_cols and node_levels arguments.
Requires ggsankey (install from GitHub):
remotes::install_github("davidsjoberg/ggsankey")
SAS/R workflow → R equivalent
sid_dta$C2 <- factor(...) with gr2_names ordering |
Factor levels set by node_levels argument |
make_long(C2, …, C9) |
.make_sankey_long() (internal) |
geom_sankey() + geom_sankey_label()
|
hv_sankey() + plot()
|
brewer.pal(9, "Set1")[c(2,6,8,4,3,5,7,1,9)] |
Default node_colours
|
To reproduce the original analysis with patient-level cluster assignments:
# Build a data frame with one row per patient, columns C2..C9 (factor)
grp_dta <- data.frame(
C2 = factor(pm2clusters_labels, levels = gr2_names),
C3 = factor(pm3clusters_labels, levels = gr3_names),
# ... etc.
C9 = factor(pm9clusters_labels, levels = gr9_names)
)
sk_grp <- hv_sankey(
grp_dta,
cluster_cols = paste0("C", 2:9),
node_levels = gr9_names
)
plot(sk_grp) +
labs(x = NULL) +
hv_theme("poster")
UpSet plot (tp.complexUpset.R)
Port: tp.complexUpset.R
R equivalent: hv_upset()
Spaghetti / individual trajectories (tp.dp.spaghetti.echo.R)
Port: tp.dp.spaghetti.echo.R
R equivalent: hv_spaghetti()
The template covers nine figures across three echo outcomes (AV mean gradient, AV area, DVI) in unstratified and sex-stratified variants, plus an ordinal MV regurgitation grade plot (plot_9).
Unstratified — AV mean gradient full range (plot_1)
Unstratified — zoomed y-axis (plot_3)
Stratified by sex (plot_2 / plot_4)
Template uses values=c("red", "blue"); modernised equivalents below.
plot(sp_col) +
scale_colour_manual(
values = c(Female = "firebrick", Male = "steelblue"),
name = NULL
) +
scale_x_continuous(breaks = seq(0, 5, 1)) +
scale_y_continuous(breaks = seq(0, 80, 20)) +
coord_cartesian(xlim = c(0, 5), ylim = c(0, 80)) +
labs(x = "Years", y = "AV Mean Gradient (mmHg)") +
hv_theme("poster")
AV area y-scale (plot_5 / plot_6)
plot(sp_col) +
scale_colour_manual(
values = c(Female = "firebrick", Male = "steelblue"),
name = NULL
) +
scale_x_continuous(breaks = seq(0, 5, 1)) +
scale_y_continuous(breaks = seq(0, 5, 1)) +
coord_cartesian(xlim = c(0, 5), ylim = c(0, 5)) +
labs(x = "Years", y = "AV Area (EOA) (cm\u00b2)") +
hv_theme("poster")
DVI y-scale (plot_7 / plot_8)
plot(sp_col) +
scale_colour_manual(
values = c(Female = "firebrick", Male = "steelblue"),
name = NULL
) +
scale_x_continuous(breaks = seq(0, 5, 1)) +
scale_y_continuous(breaks = seq(0, 1.25, 0.25)) +
coord_cartesian(xlim = c(0, 5), ylim = c(0, 1.25)) +
labs(x = "Years", y = "DVI") +
hv_theme("poster")
Ordinal y-axis — MV regurgitation grade (plot_9)
dta_ord <- dta
dta_ord$value <- round(pmin(3, pmax(0, dta$value / 12)))
levels(dta_ord$group) <- c("Early", "Late")
sp_ord <- hv_spaghetti(dta_ord, colour_col = "group")
plot(sp_ord, y_labels = c(None = 0, Mild = 1, Moderate = 2, Severe = 3)) +
scale_colour_manual(
values = c(Early = "steelblue", Late = "red2"),
name = NULL
) +
scale_x_continuous(breaks = seq(0, 6, 1)) +
coord_cartesian(xlim = c(0, 6), ylim = c(0, 3)) +
labs(x = "Years after Procedure", y = "MV Regurgitation Grade") +
hv_theme("poster")
Trends over time (tp.*.trends.*)
Ports: tp.rp.trends.sas, tp.lp.trends.sas, tp.lp.trends.age.sas, tp.lp.trends.polytomous.sas, tp.dp.trends.R
R equivalent: hv_trends()
tp.rp.trends.sas — cases/year and age (1968–2000 by 4)
one <- sample_trends_data(n = 600, year_range = c(1968L, 2000L), groups = NULL)
tr_one <- hv_trends(one, group_col = NULL)
# Cases/year: axisy order=(0 to 10 by 2)
plot(tr_one) +
scale_x_continuous(limits = c(1968, 2000), breaks = seq(1968, 2000, 4)) +
scale_y_continuous(limits = c(0, 10), breaks = seq(0, 10, 2)) +
labs(x = "Year", y = "Cases/year") +
hv_theme("poster")
# Age: axisy order=(30 to 70 by 10)
plot(tr_one) +
scale_x_continuous(limits = c(1968, 2000), breaks = seq(1968, 2000, 4)) +
scale_y_continuous(limits = c(30, 70), breaks = seq(30, 70, 10)) +
labs(x = "Year", y = "Age (years)") +
hv_theme("poster")
tp.lp.trends.sas — binary % outcomes (1970–2000 by 10, y 0–100 by 10)
Multiple outcomes on one figure. CGM version: axisx order=(1970 to 2000 by 10), axisy order=(0 to 100 by 20).
dta_lp <- sample_trends_data(
n = 800, year_range = c(1970L, 2000L),
groups = c("Shock %", "Pre-op IABP %", "Inotropes %"))
tr_lp <- hv_trends(dta_lp)
plot(tr_lp) +
scale_colour_manual(
values = c("Shock %" = "steelblue", "Pre-op IABP %" = "firebrick",
"Inotropes %" = "forestgreen"), name = NULL) +
scale_shape_manual(
values = c("Shock %" = 16L, "Pre-op IABP %" = 15L, "Inotropes %" = 17L),
name = NULL) +
scale_x_continuous(limits = c(1970, 2000), breaks = seq(1970, 2000, 10)) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 10)) +
coord_cartesian(xlim = c(1970, 2000), ylim = c(0, 100)) +
labs(x = "Year", y = "Percent (%)") +
hv_theme("poster")
tp.lp.trends.age.sas — age on x-axis (25–85 by 10, y 0–100 by 20)
dta_age <- sample_trends_data(
n = 600, year_range = c(25L, 85L),
groups = c("Repair %", "Bioprosthesis %"), seed = 7L)
tr_age <- hv_trends(dta_age)
plot(tr_age) +
scale_colour_manual(
values = c("Repair %" = "steelblue", "Bioprosthesis %" = "firebrick"),
name = NULL) +
scale_x_continuous(limits = c(25, 85), breaks = seq(25, 85, 10)) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 20)) +
coord_cartesian(xlim = c(25, 85), ylim = c(0, 100)) +
labs(x = "Age (years)", y = "Percent (%)") +
hv_theme("poster")
tp.lp.trends.polytomous.sas — repair types (1990–1999 by 1, y 0–100 by 10)
dta_poly <- sample_trends_data(
n = 800, year_range = c(1990L, 1999L),
groups = c("CE", "Cosgrove", "Periguard", "DeVega"), seed = 5L)
tr_poly <- hv_trends(dta_poly)
plot(tr_poly) +
scale_colour_manual(
values = c(CE = "steelblue", Cosgrove = "firebrick",
Periguard = "forestgreen", DeVega = "goldenrod3"),
name = "Repair type") +
scale_shape_manual(
values = c(CE = 15L, Cosgrove = 19L, Periguard = 17L, DeVega = 18L),
name = "Repair type") +
scale_x_continuous(limits = c(1990, 1999), breaks = seq(1990, 1999, 1)) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 10)) +
coord_cartesian(xlim = c(1990, 1999), ylim = c(0, 100)) +
labs(x = "Year", y = "Percent (%)") +
hv_theme("poster")
tp.dp.trends.R — LV mass index (1995–2015 by 5, y 0–200 by 50)
dta_lv <- sample_trends_data(n = 800, year_range = c(1995L, 2015L),
groups = NULL, seed = 3L)
tr_lv <- hv_trends(dta_lv, group_col = NULL)
plot(tr_lv) +
scale_x_continuous(limits = c(1995, 2015), breaks = seq(1995, 2015, 5)) +
scale_y_continuous(limits = c(0, 200), breaks = seq(0, 200, 50)) +
coord_cartesian(xlim = c(1995, 2015), ylim = c(0, 200)) +
labs(x = "Years", y = "LV Mass Index") +
hv_theme("poster")
tp.dp.trends.R — hospital LOS with annotation (1985–2015 by 5, y 0–20 by 5)
dta_los <- sample_trends_data(n = 800, year_range = c(1985L, 2015L),
groups = NULL, seed = 11L)
tr_los <- hv_trends(dta_los, group_col = NULL)
plot(tr_los) +
scale_x_continuous(limits = c(1985, 2015), breaks = seq(1985, 2015, 5)) +
scale_y_continuous(limits = c(0, 20), breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(1985, 2015), ylim = c(0, 20)) +
annotate("text", x = 1995, y = 18,
label = "Trend: Hospital Length of Stay", size = 4.5) +
labs(x = "Years", y = "Hospital LOS (Days)") +
hv_theme("poster")
Longitudinal patient counts (tp.dp.longitudinal_patients_measures.R)
Port: tp.dp.longitudinal_patients_measures.R
R equivalent: hv_longitudinal()
Mirror histogram (propensity score)
R equivalents: tp.lp.mirror-histogram_SAVR-TF-TAVR.R (binary-match), tp.lp.mirror_histo_before_after_wt.R (weighted IPTW)
Both scripts are superseded by hv_mirror_hist(). Call plot(mh) to render the histogram. Diagnostics are accessible via mh$tables$diagnostics; working data via mh$tables$working. Compose scales, annotations, and theme with the usual + operator.
Binary-match mode (tp.lp.mirror-histogram_SAVR-TF-TAVR.R)
dta <- sample_mirror_histogram_data(n = 400, separation = 1.5)
mh <- hv_mirror_hist(dta) # defaults: prob_t / tavr / match
plot(mh) +
scale_fill_manual(
values = c(
before_g0 = "white", matched_g0 = "green1",
before_g1 = "white", matched_g1 = "green4"
),
guide = "none"
) +
scale_x_continuous(limits = c(0, 100), breaks = seq(0, 100, 10)) +
annotate("text", x = 20, y = 100, label = "SAVR", size = 7) +
annotate("text", x = 20, y = -100, label = "TF-TAVR", size = 7) +
labs(x = "Propensity Score (%)", y = "Number of Patients") +
hv_theme("poster")
Weighted IPTW mode (tp.lp.mirror_histo_before_after_wt.R)
dta <- sample_mirror_histogram_data(n = 400, add_weights = TRUE)
mh_wt <- hv_mirror_hist(
dta,
group_labels = c("Limited", "Extended"),
weight_col = "mt_wt"
)
plot(mh_wt) +
scale_fill_manual(
values = c(
before_g0 = "white", weighted_g0 = "blue",
before_g1 = "white", weighted_g1 = "red"
),
guide = "none"
) +
scale_x_continuous(limits = c(0, 100), breaks = seq(0, 100, 10)) +
annotate("text", x = 30, y = 150, label = "Limited", color = "blue", size = 5) +
annotate("text", x = 30, y = -70, label = "Extended", color = "red", size = 5) +
labs(x = "Propensity Score (%)", y = "#") +
hv_theme("poster")
Using themes
All plot() calls return an unstyled ggplot object. Add a theme as the final layer:
| Dark PowerPoint slide |
hv_theme("dark_ppt") |
device=ppt |
| Light PowerPoint slide |
hv_theme("light_ppt") |
device=ppt with white background |
| Journal manuscript |
hv_theme("manuscript") |
device=eps / PDF |
| Conference poster |
hv_theme("poster") |
Large-font poster |
You can pass base_size to any theme to scale all text simultaneously:
p + hv_theme("manuscript", base_size = 10) # smaller text for double-column
p + hv_theme("poster", base_size = 24) # larger text for A0 poster
PowerPoint
save_ppt() inserts ggplot objects into a PowerPoint file as editable DrawingML vector graphics via the officer and rvg packages — shapes, lines, and text remain individually selectable in PowerPoint after export. The first argument is object (not plot); the output path is powerpoint (not file or filename).
# Locate the bundled Cleveland Clinic slide template
template <- system.file("ClevelandClinic.pptx", package = "hvtiPlotR")
# Single slide — apply a PPT theme before saving
p_ppt <- p +
scale_colour_manual(values = c("steelblue"), guide = "none") +
scale_fill_manual(values = c("steelblue"), guide = "none") +
labs(x = "Years", y = "Prevalence (%)") +
hv_theme("dark_ppt")
save_ppt(
object = p_ppt,
template = template,
powerpoint = "figures/afib_prevalence.pptx",
slide_titles = "AF Prevalence over Time"
)
# Multiple plots — one slide per figure in a single call
save_ppt(
object = list(fig1 = p_binary, fig2 = p_multi),
template = template,
powerpoint = "figures/np_curves.pptx",
slide_titles = c("Binary Outcome", "Multi-group Comparison")
)
PDF / TIFF for journals
ggsave("figures/afib_prevalence.pdf", p, width = 3.5, height = 3.5, units = "in")
ggsave("figures/afib_prevalence.tiff", p, width = 3.5, height = 3.5,
units = "in", dpi = 600)
Parametric hazard/survival (tp.hs.*)
Ports: tp.hs.dead.setup.sas, tp.hs.dead_uses_setup.sas, tp.hs.dead.procedure.tdepth.sas, tp.hs.dead.conditional.setup.sas, tp.hs.dead.conditional.uses_setup.sas, tp.hs.dead.compare_benefit.setup.sas, tp.hs.uslife_estimates_generate_stratify_.age.sas, tp.hs.uslife_generates_matched_estimates.sas
R equivalents: hazard_plot(), survival_difference_plot()
The tp.hs.* family extends tp.hp.dead.* by using the %hazpred macro to generate patient-specific parametric survival predictions from a fitted multivariable hazard model. The setup templates compute and store:
-
Cumulative hazard per patient at last follow-up (for observed vs. expected goodness-of-fit tests via
%chisqgf).
-
Individual survivorship curves on a fine time grid (for mean-curve aggregation with
PROC SUMMARY).
Subsequent use templates stratify the aggregated mean curves by a covariate and overlay Kaplan-Meier estimates from %kaplan. The resulting SAS datasets map directly to hazard_plot() in R:
Parametric survival with KM overlay
Reproduces the core figure from tp.hs.dead_uses_setup.sas and tp.hs.dead.procedure.tdepth.sas: a mean parametric survival curve (solid line with confidence band) overlaid with Kaplan-Meier empirical estimates (symbols with error bars).
dat_hp <- sample_hazard_data(n = 500, time_max = 10)
emp_hp <- sample_hazard_empirical(n = 500, time_max = 10, n_bins = 6)
hazard_plot(
dat_hp,
estimate_col = "survival",
lower_col = "surv_lower",
upper_col = "surv_upper",
empirical = emp_hp,
emp_lower_col = "lower",
emp_upper_col = "upper"
) +
scale_colour_manual(values = c("steelblue"), guide = "none") +
scale_fill_manual(values = c("steelblue"), guide = "none") +
scale_x_continuous(limits = c(0, 10), breaks = 0:10) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 10),
labels = function(x) paste0(x, "%")) +
labs(x = "Years after Operation", y = "Survival (%)") +
hv_theme("poster")
Stratified by covariate
tp.hs.dead.procedure.tdepth.sas overlays curves for depth-of-invasion groups (tdepth = 1, 2, 3). Pass group_col to hazard_plot() and supply matched sample_hazard_data() / sample_hazard_empirical() calls with a groups vector whose names become the legend labels.
dat_strat <- sample_hazard_data(
n = 500, time_max = 10,
groups = c("pT1" = 0.4, "pT2" = 1.0, "pT3" = 1.8)
)
emp_strat <- sample_hazard_empirical(
n = 500, time_max = 10, n_bins = 5,
groups = c("pT1" = 0.4, "pT2" = 1.0, "pT3" = 1.8)
)
hazard_plot(
dat_strat,
estimate_col = "survival",
lower_col = "surv_lower",
upper_col = "surv_upper",
group_col = "group",
empirical = emp_strat,
emp_lower_col = "lower",
emp_upper_col = "upper"
) +
scale_colour_manual(
values = c("pT1" = "steelblue", "pT2" = "forestgreen", "pT3" = "firebrick"),
name = NULL
) +
scale_fill_manual(
values = c("pT1" = "steelblue", "pT2" = "forestgreen", "pT3" = "firebrick"),
guide = "none"
) +
scale_x_continuous(limits = c(0, 10), breaks = 0:10) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 10),
labels = function(x) paste0(x, "%")) +
labs(x = "Years after Operation", y = "Survival (%)") +
hv_theme("poster")
Conditional survival after hospital discharge
tp.hs.dead.conditional.setup.sas computes individual survivorship curves starting from the time of hospital discharge rather than the operation date. The conditional survival S(t | discharge) is S(t) / S(t_discharge) for each patient. In R, the same hazard_plot() call is used; only the x-axis label and data preparation differ.
dat_cond <- sample_hazard_data(n = 500, time_max = 10)
emp_cond <- sample_hazard_empirical(n = 500, time_max = 10, n_bins = 6)
hazard_plot(
dat_cond,
estimate_col = "survival",
lower_col = "surv_lower",
upper_col = "surv_upper",
empirical = emp_cond,
emp_lower_col = "lower",
emp_upper_col = "upper"
) +
scale_colour_manual(values = c("steelblue"), guide = "none") +
scale_fill_manual(values = c("steelblue"), guide = "none") +
scale_x_continuous(limits = c(0, 10), breaks = 0:10) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 10),
labels = function(x) paste0(x, "%")) +
labs(x = "Years after Discharge", y = "Survival after Discharge (%)") +
hv_theme("poster")
US life-table overlay
tp.hs.uslife_* templates call %usmatchd to generate age-, sex-, and race-matched US population life-table survival curves for each patient. sample_life_table() provides representative matched curves stratified by age group for use in R examples.
dat_age <- sample_hazard_data(
n = 600, time_max = 10,
groups = c("<60" = 0.4, "60\u201385" = 1.0, "\u226585" = 2.0)
)
emp_age <- sample_hazard_empirical(
n = 600, time_max = 10, n_bins = 6,
groups = c("<60" = 0.4, "60\u201385" = 1.0, "\u226585" = 2.0)
)
lt <- sample_life_table(
age_groups = c("<60", "60\u201385", "\u226585"),
age_mids = c(50, 72, 88),
time_max = 10
)
hazard_plot(
dat_age,
estimate_col = "survival",
lower_col = "surv_lower",
upper_col = "surv_upper",
group_col = "group",
empirical = emp_age,
emp_lower_col = "lower",
emp_upper_col = "upper",
reference = lt,
ref_estimate_col = "survival",
ref_group_col = "group"
) +
scale_colour_manual(
values = c("<60" = "steelblue", "60\u201385" = "forestgreen",
"\u226585" = "firebrick"),
name = "Age group"
) +
scale_fill_manual(
values = c("<60" = "steelblue", "60\u201385" = "forestgreen",
"\u226585" = "firebrick"),
guide = "none"
) +
scale_x_continuous(limits = c(0, 10), breaks = 0:10) +
scale_y_continuous(limits = c(0, 100), breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
labs(x = "Years after Operation", y = "Survival (%)",
caption = "Dashed lines: US population life table") +
hv_theme("poster")
Treatment benefit distribution
tp.hs.dead.compare_benefit.setup.sas computes, for each patient, the difference in predicted survival at a fixed time point (5 years) under two treatment arms (e.g. ASA vs. no ASA). The distribution of these individual differences shows who benefits and by how much. Use survival_difference_plot() to plot the mean curve over time.
diff_dat <- sample_survival_difference_data(
n = 500,
groups = c("No ASA" = 1.0, "ASA" = 0.75)
)
survival_difference_plot(
diff_dat,
lower_col = "diff_lower",
upper_col = "diff_upper"
) +
scale_colour_manual(values = c("steelblue"), guide = "none") +
scale_fill_manual(values = c("steelblue"), guide = "none") +
ggplot2::geom_hline(yintercept = 0, linetype = "dashed",
colour = "grey50") +
scale_x_continuous(limits = c(0, 10), breaks = 0:10) +
scale_y_continuous(limits = c(-5, 40),
labels = function(x) paste0(x, "%")) +
labs(x = "Years", y = "Survival Difference (%)") +
hv_theme("poster")
Adding new ports
As additional SAS templates are ported to R, update this guide:
Add a row to the lookup table at the top of this file with the SAS template name, family prefix, R function, and section anchor.
Add a section following the existing pattern: name the port, describe the SAS workflow, show the R equivalent with a runnable example, and document any column name mapping differences.
Export the new R function by adding @export to its roxygen block and running devtools::document() to regenerate NAMESPACE.
Update DESCRIPTION if a new package dependency is required.
Templates currently planned for porting:
-
tp.ce.states.* competing-events / state-occupancy → (pending)
-
tp.gp.* grouped longitudinal ordinal models → (pending)
Session info
R version 4.5.3 (2026-03-11)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ggplot2_4.0.2 hvtiPlotR_2.0.0.9010
loaded via a namespace (and not attached):
[1] generics_0.1.4 tidyr_1.3.2 fontLiberation_0.1.0
[4] xml2_1.5.2 lattice_0.22-9 digest_0.6.39
[7] magrittr_2.0.4 evaluate_1.0.5 grid_4.5.3
[10] RColorBrewer_1.1-3 fastmap_1.2.0 Matrix_1.7-4
[13] jsonlite_2.0.0 zip_2.3.3 survival_3.8-6
[16] purrr_1.2.1 scales_1.4.0 fontBitstreamVera_0.1.1
[19] textshaping_1.0.5 cli_3.6.5 rlang_1.1.7
[22] fontquiver_0.2.1 splines_4.5.3 withr_3.0.2
[25] yaml_2.3.12 otel_0.2.0 gdtools_0.5.0
[28] tools_4.5.3 officer_0.7.3 uuid_1.2-2
[31] dplyr_1.2.0 colorspace_2.1-2 ComplexUpset_1.3.3
[34] vctrs_0.7.2 R6_2.6.1 lifecycle_1.0.5
[37] ragg_1.5.2 pkgconfig_2.0.3 pillar_1.11.1
[40] gtable_0.3.6 glue_1.8.0 Rcpp_1.1.1
[43] systemfonts_1.3.2 xfun_0.57 rvg_0.4.1
[46] tibble_3.3.1 tidyselect_1.2.1 knitr_1.51
[49] farver_2.1.2 htmltools_0.5.9 patchwork_1.3.2
[52] labeling_0.4.3 rmarkdown_2.31 ggalluvial_0.12.6
[55] compiler_4.5.3 S7_0.2.1 askpass_1.2.1
[58] openssl_2.3.5