The Composition Pattern
Every hvtiPlotR plot is built in two steps: a constructor (hv_*()) that shapes the data, followed by plot() that renders a bare ggplot object. No colour scales, axis labels, or theme are applied by either step. Decoration is added by chaining layers with +:
plot(hv_*(...)) +
scale_colour_*() + # data colours
scale_fill_*() + # fill colours
labs() + # axis labels, title, caption
annotate() + # text/arrows placed on the panel
coord_cartesian() + # viewport cropping
hv_theme() # non-data formattingThis vignette demonstrates each decorator in turn, using hv_trends() and hv_survival() as representative base plots.
# Trends data — multi-group continuous outcome over time
dta_trends <- sample_trends_data(n = 600, seed = 42)
p_base <- plot(hv_trends(dta_trends))
# KM data — survival curve
dta_km <- sample_survival_data(n = 500, seed = 42)
km <- hv_survival(dta_km)Themes
The hvtiPlotR package provides four themes via hv_theme(style). The style argument selects the output target.
| Style | Target |
|---|---|
"manuscript" |
Journal PDF, black-on-white |
"poster" |
Conference poster, slightly larger text |
"light_ppt" |
PowerPoint on white/light background |
"dark_ppt" |
PowerPoint on dark/blue background |
Manuscript
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("manuscript")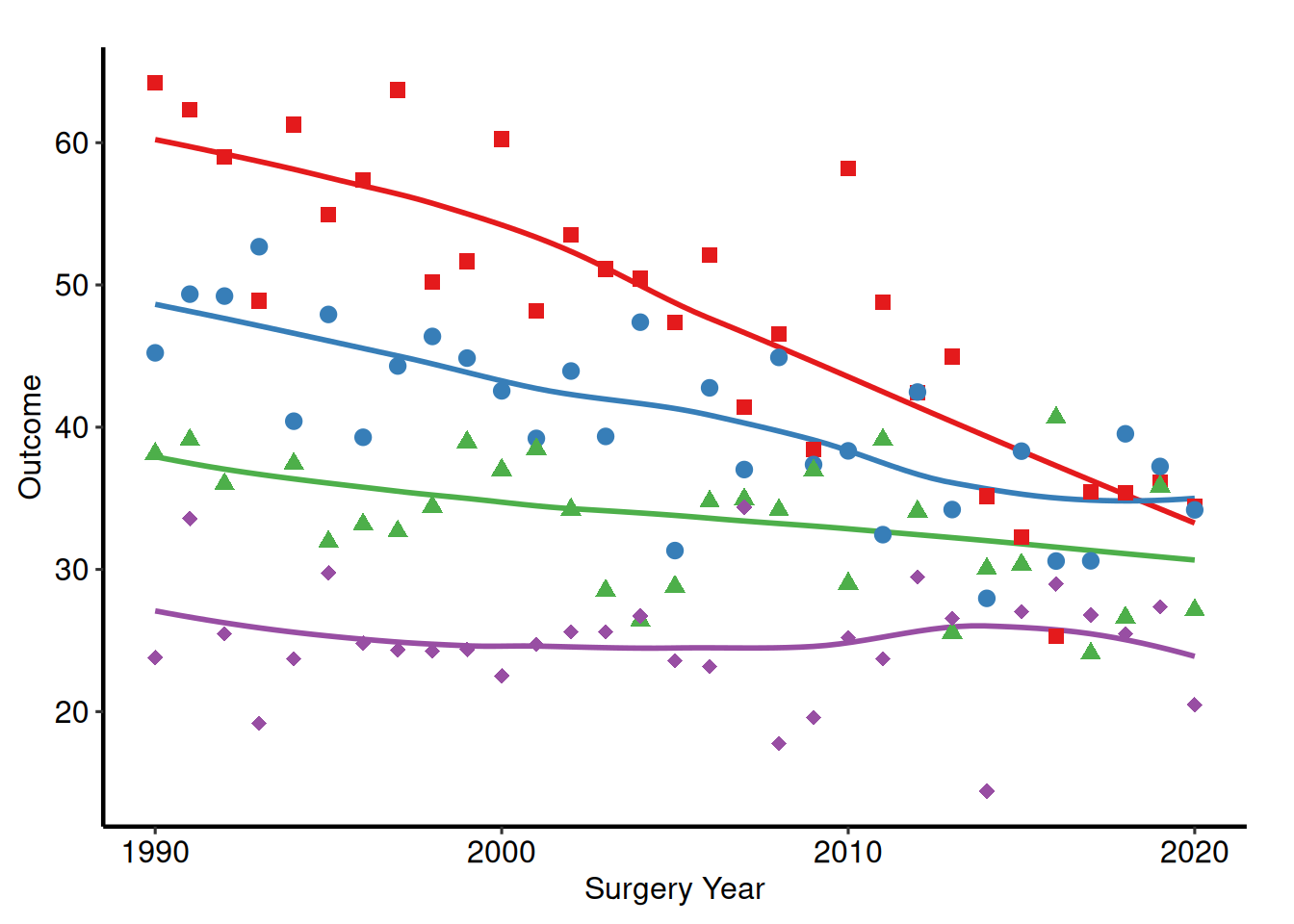
Poster
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster")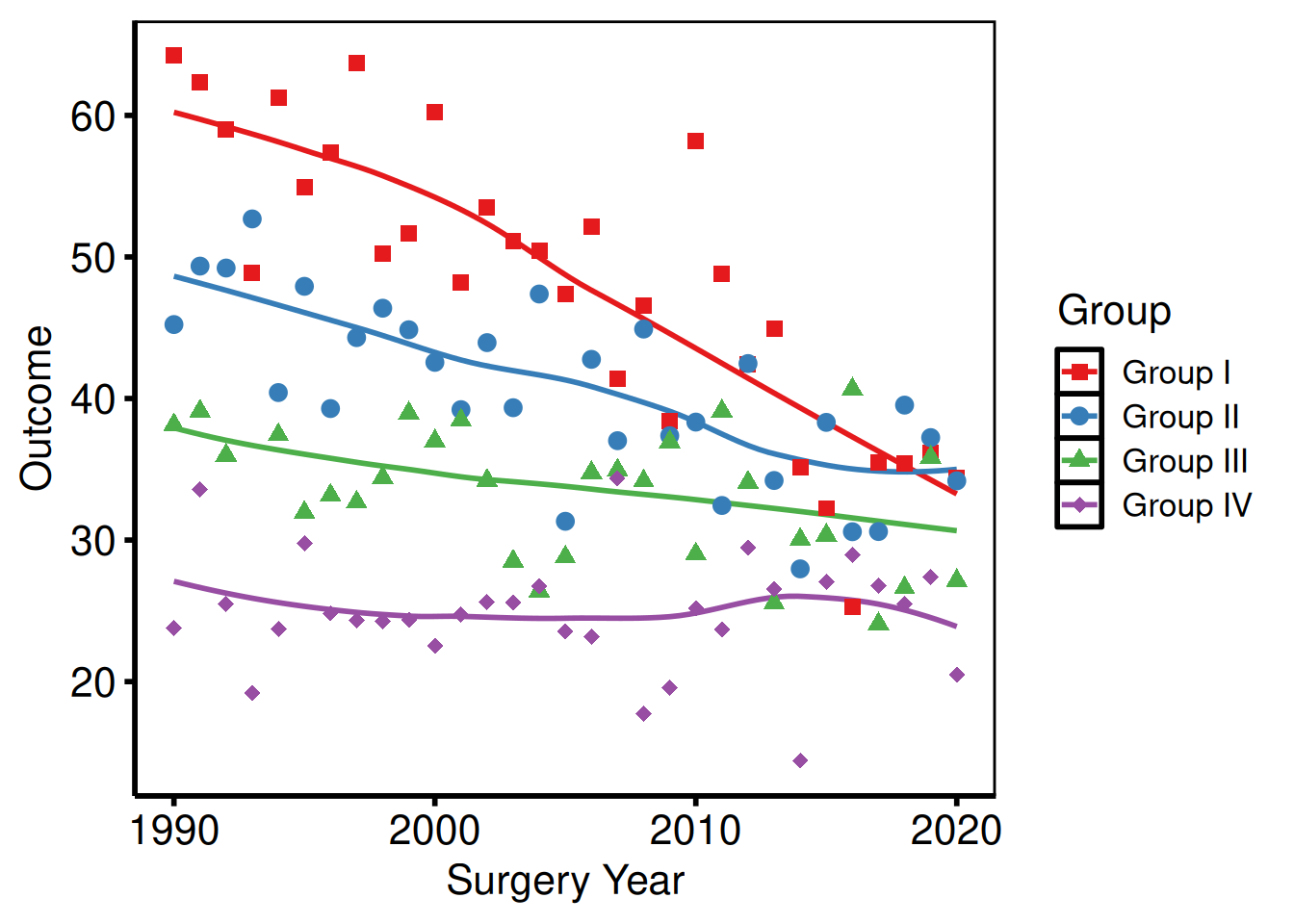
Light PowerPoint
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("light_ppt")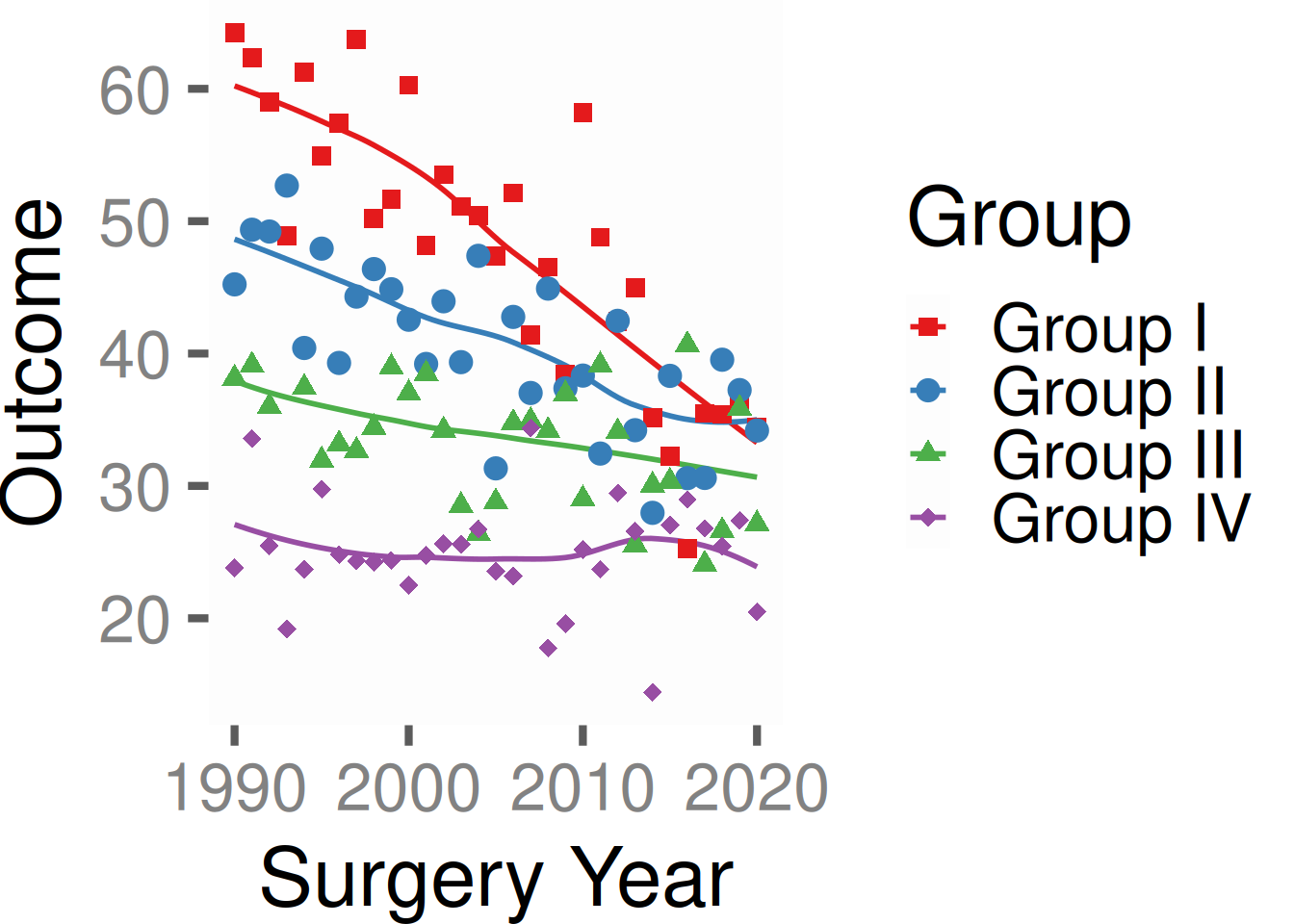
Dark PowerPoint
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("dark_ppt") +
theme(plot.background = element_rect(fill = "navy", colour = "navy"))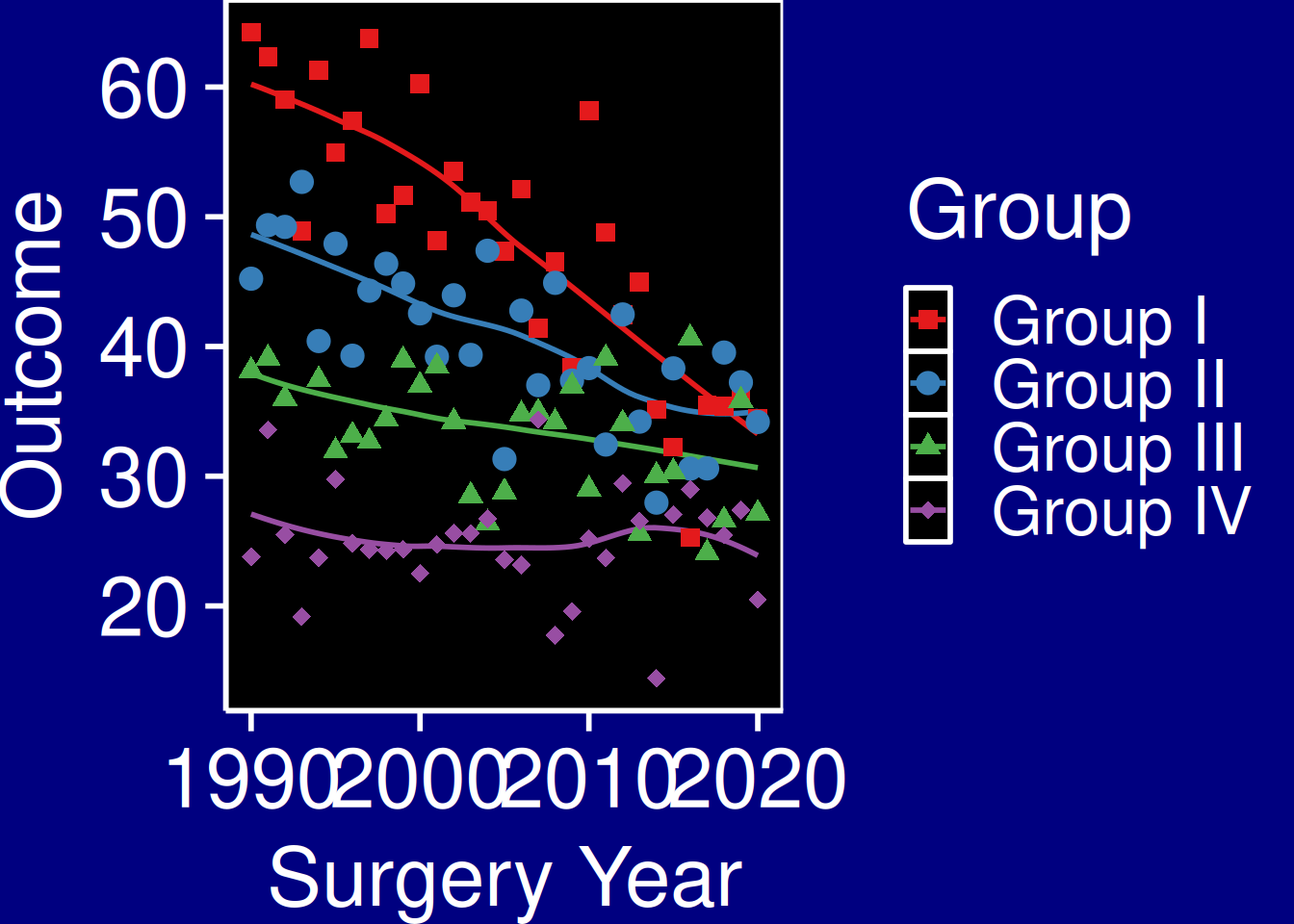
Colour Scales
scale_colour_* controls line and point colours; scale_fill_* controls filled areas (ribbons, bars). Both share the same name (legend title) and guide (legend display) arguments.
Manual colours
Use scale_colour_manual() when assigning specific brand or convention colours to known levels.
plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(x = "Years after Operation", y = "Freedom from Death (%)") +
hv_theme("poster")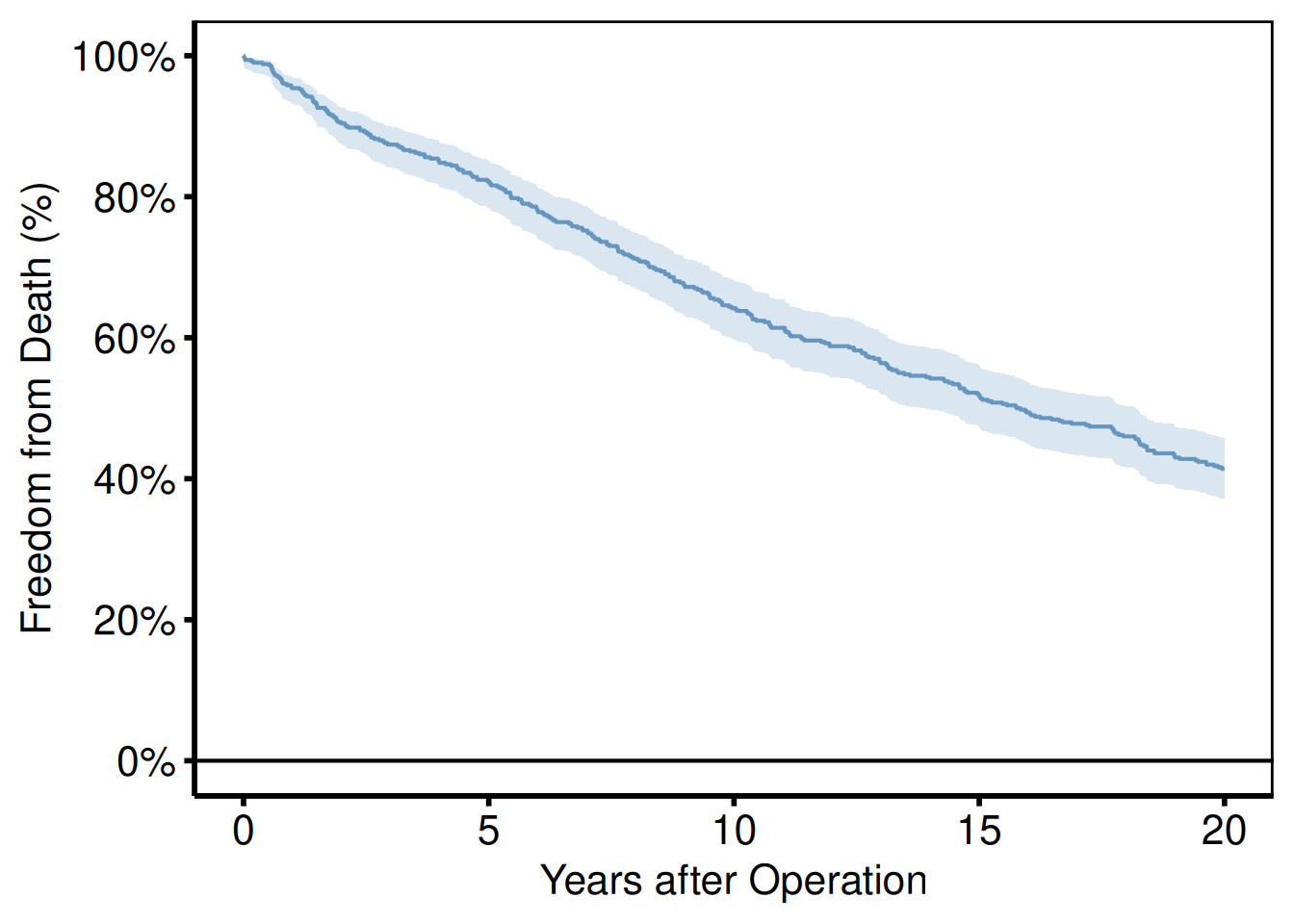
ColorBrewer palettes
scale_colour_brewer() applies a ColorBrewer palette — safe, perceptually uniform, and print-friendly. Use palette = "Set1" for categorical data, "RdYlGn" for diverging, "Blues" for sequential.
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster")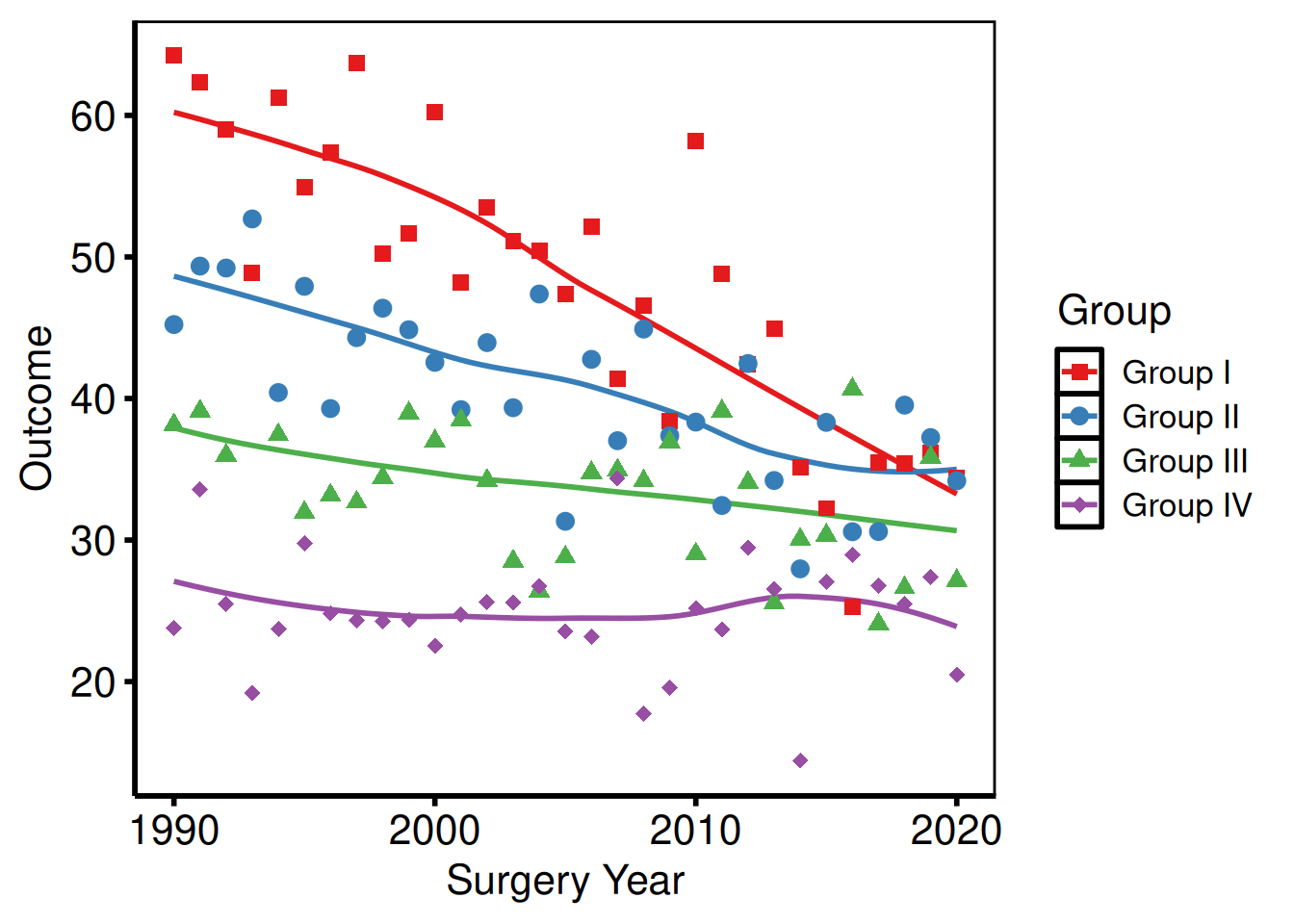
Suppressing legends
Pass guide = "none" to any scale to remove its legend entry. Use this when colour is self-evident from axis labels or annotations.
p_base +
scale_colour_brewer(palette = "Dark2", guide = "none") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
guide = "none"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster")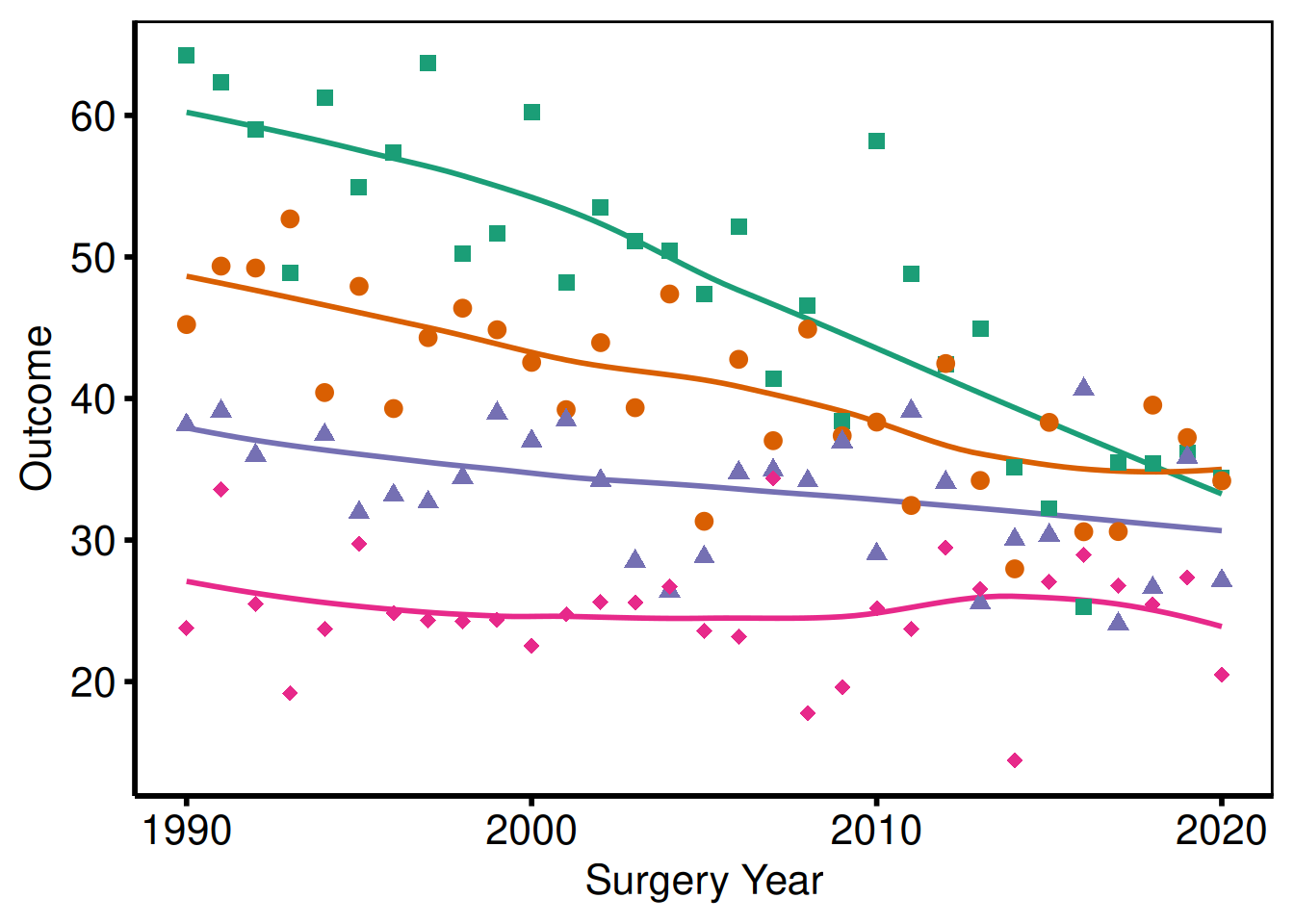
Labels and Annotations
labs()
labs() sets the axis, legend, title, subtitle, and caption text. Set axis labels here rather than inside the plot function so they can be overridden per project.
plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(
title = "Overall Survival",
x = "Years after Operation",
y = "Freedom from Death (%)",
caption = "Logit CI, \u03b1 = 0.6827 (1 SD)"
) +
hv_theme("poster")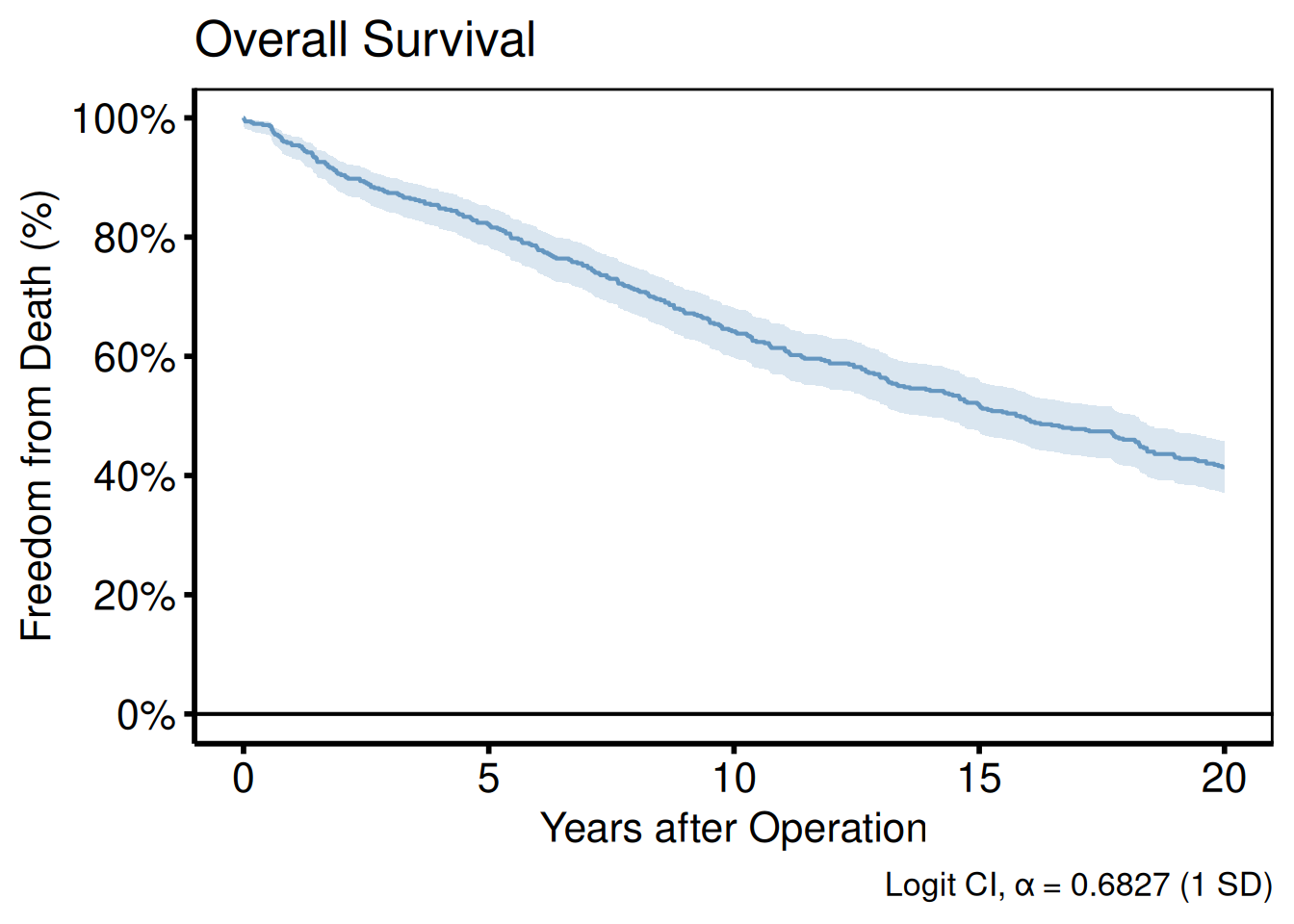
annotate()
annotate() places text, segments, or rectangles at fixed data coordinates. Use it for sample size callouts, phase labels, or directional arrows.
plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(x = "Years after Operation", y = "Freedom from Death (%)") +
annotate("text", x = 1, y = 5,
label = paste0("n = ", nrow(dta_km)),
hjust = 0, size = 3.5) +
annotate("segment", x = 10, xend = 10, y = 30, yend = 50,
arrow = arrow(length = unit(0.2, "cm")), colour = "grey40") +
annotate("text", x = 10.3, y = 40,
label = "Median survival", hjust = 0, size = 3, colour = "grey40") +
hv_theme("poster")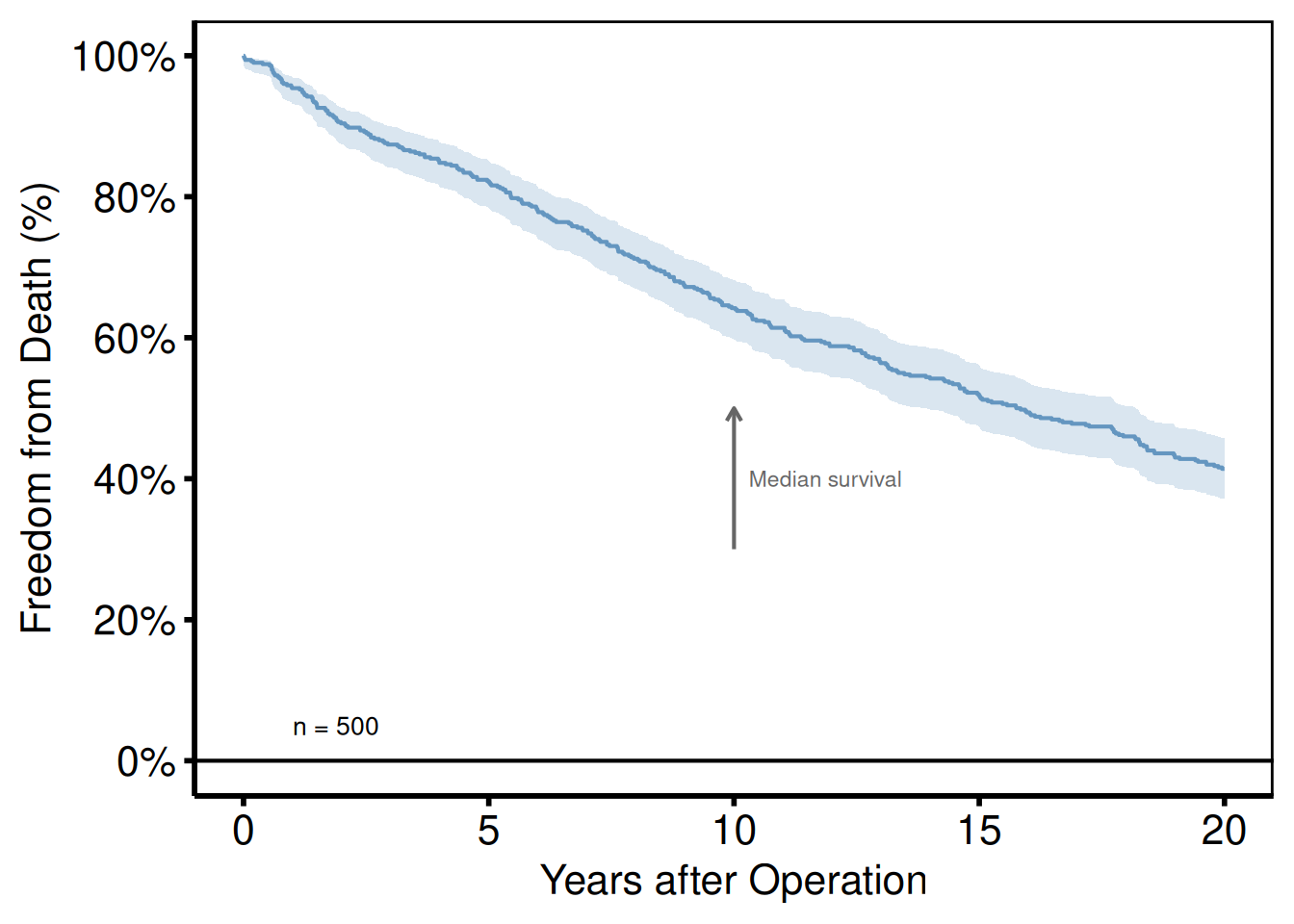
coord_cartesian()
coord_cartesian() crops the viewport without dropping data, preserving LOESS fits computed on the full range.
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
coord_cartesian(xlim = c(1995, 2020), ylim = c(20, 70)) +
hv_theme("poster")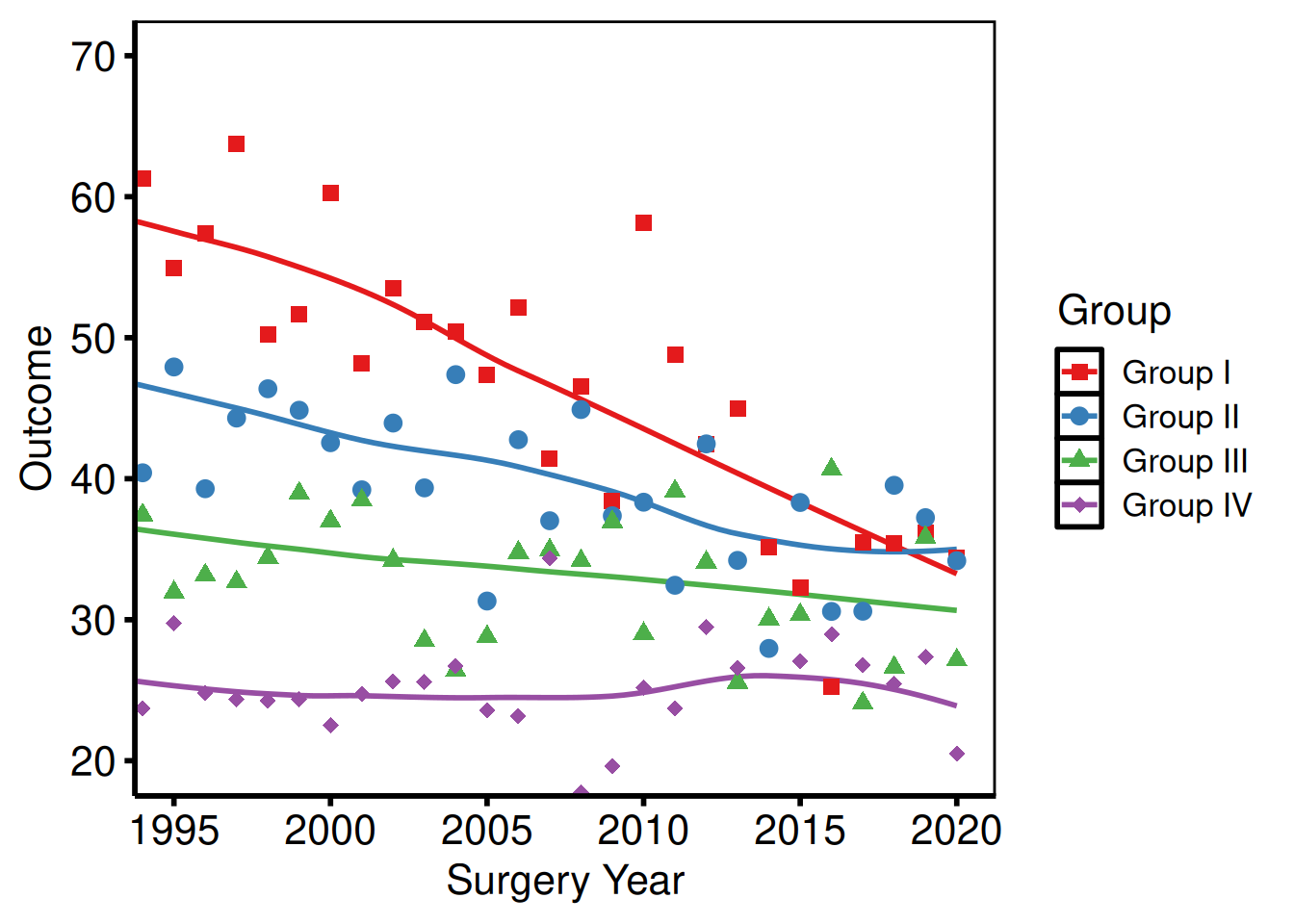
Saving Figures
Manuscript PDF
Use ggsave() with width = 11, height = 8.5 (US Letter landscape) for manuscript figures. Assign the fully composed plot to a variable first so the same object is both displayed in the session and written to disk.
p_ms <- p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome (%)") +
hv_theme("manuscript")
ggsave(
filename = "../graphs/trends_manuscript.pdf",
plot = p_ms,
width = 11,
height = 8.5
)Poster PDF
Poster figures are typically larger and use hv_theme("poster"). Adjust dimensions to match the poster panel size.
p_poster <- p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome (%)") +
hv_theme("poster")
ggsave(
filename = "../graphs/trends_poster.pdf",
plot = p_poster,
width = 14,
height = 10
)PowerPoint slides
save_ppt() inserts ggplot objects into a PowerPoint file as editable DrawingML vector graphics via the officer and rvg packages — shapes, lines, and text remain selectable in PowerPoint after export.
Key arguments:
| Argument | Default | Notes |
|---|---|---|
object |
— | A single ggplot or a named/unnamed list of ggplots |
template |
"../graphs/RD.pptx" |
Existing .pptx used as the slide template |
powerpoint |
"../graphs/pptExample.pptx" |
Output file path |
slide_titles |
"Plot" |
Character vector recycled to the number of plots |
layout |
"Title and Content" |
Slide layout from the template |
width / height
|
10.1 / 5.8
|
Plot area in inches |
left / top
|
0.0 / 1.2
|
Position from slide edges, in inches |
Apply hv_theme("dark_ppt") or hv_theme("light_ppt") before saving to match the slide background.
Single slide
template <- system.file("ClevelandClinic.pptx", package = "hvtiPlotR")
p_ppt <- p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome (%)") +
hv_theme("dark_ppt")
save_ppt(
object = p_ppt,
template = template,
powerpoint = here::here("graphs", "trends_slides.pptx"),
slide_titles = "Temporal Trends by Group"
)Multiple slides from a list
Pass a named list of plots and a matching vector of titles to produce one slide per plot in a single call.
dta_km2 <- sample_survival_data(n = 400, seed = 99)
km2 <- hv_survival(dta_km2)
p_km_ppt <- plot(km2) +
scale_color_manual(values = c(All = "white"), guide = "none") +
scale_fill_manual(values = c(All = "white"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(x = "Years after Operation", y = "Freedom from Death (%)") +
hv_theme("dark_ppt")
save_ppt(
object = list(trends = p_ppt, survival = p_km_ppt),
template = template,
powerpoint = here::here("graphs", "multi_slide_deck.pptx"),
slide_titles = c("Temporal Trends by Group", "Overall Survival")
)Multi-panel PDF (EDA batch output)
When generating multiple plots in a loop, gridExtra::marrangeGrob() arranges them into a grid and ggsave() writes each page.
# Build a list of plots (e.g. from an hv_eda() lapply loop)
plot_list <- lapply(
c("ef", "lv_mass", "peak_grad"),
function(yv) {
dta_eda <- sample_eda_data()
plot(hv_eda(dta_eda, x_col = "op_years", y_col = yv,
y_label = yv)) +
scale_colour_manual(values = c("steelblue"), guide = "none") +
labs(x = "Years") +
hv_theme("poster")
}
)
per_page <- 9L
for (pg in seq(1, length(plot_list), by = per_page)) {
idx <- seq(pg, min(pg + per_page - 1L, length(plot_list)))
grob <- gridExtra::marrangeGrob(plot_list[idx], nrow = 3, ncol = 3)
ggsave(
filename = sprintf(here::here("graphs", "eda_page%02d.pdf"),
ceiling(pg / per_page)),
plot = grob,
width = 14,
height = 14
)
}Legend Positioning
ggplot2 places the legend outside the panel by default. For publication figures it is often cleaner to place it inside the panel or suppress it entirely.
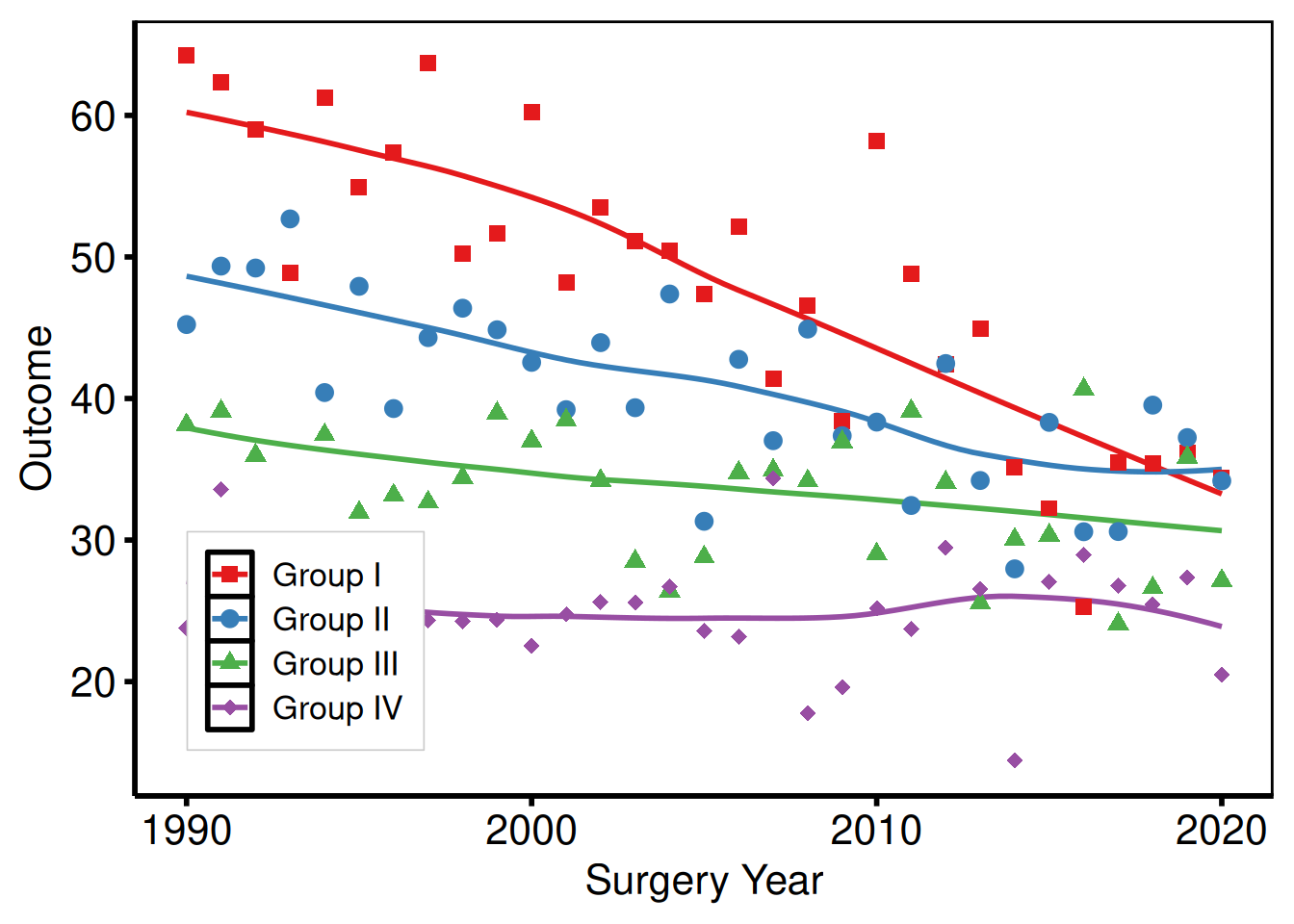
Inside the panel
Pass fractional coordinates c(x, y) to legend.position inside theme(). c(0, 0) is the bottom-left corner; c(1, 1) is the top-right.
p_base +
scale_colour_brewer(palette = "Set1", name = NULL) +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = NULL
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
legend.position = c(0.15, 0.2), # bottom-left of panel
legend.background = element_rect(fill = "white", colour = "grey80",
linewidth = 0.3)
)
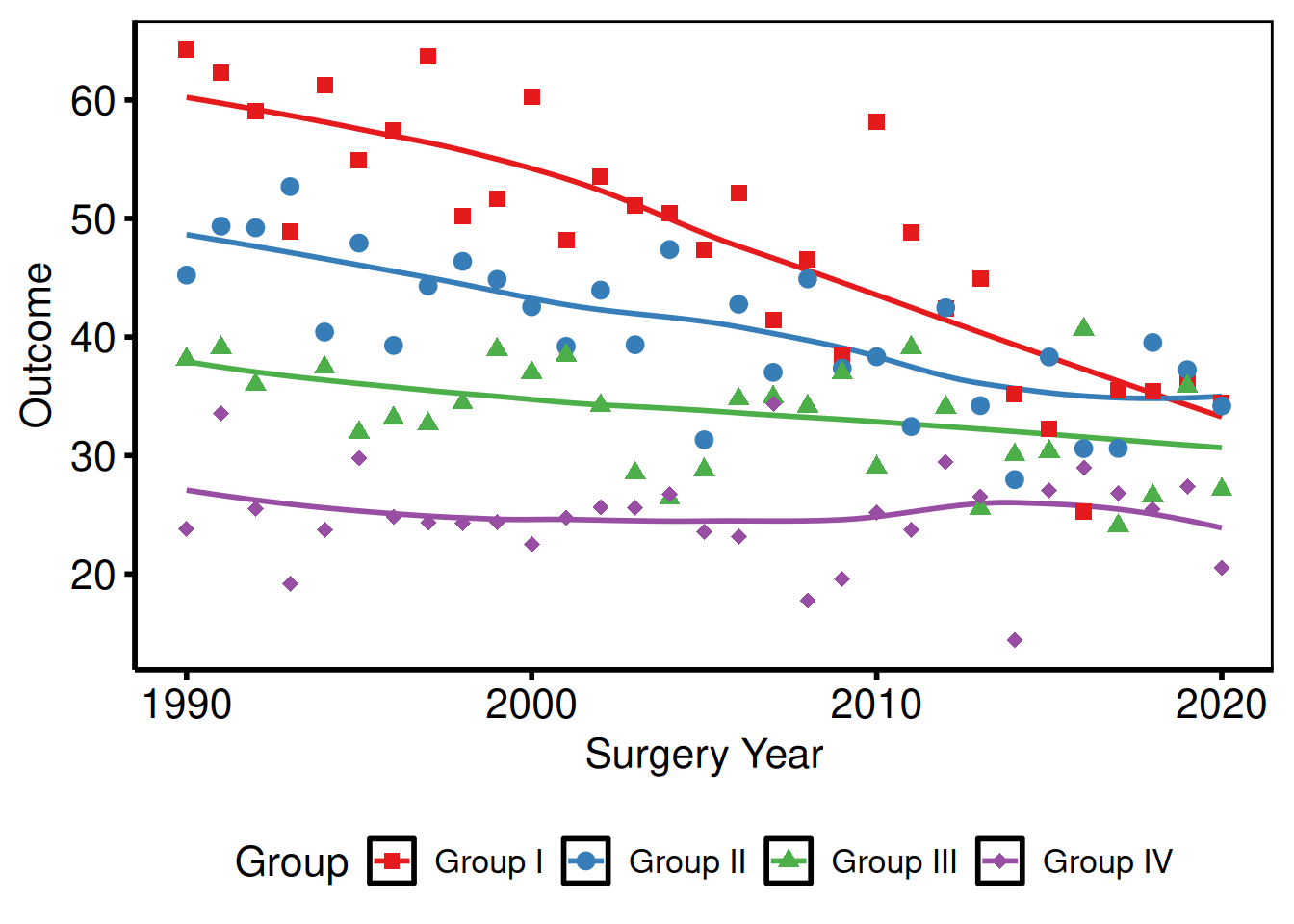
Outside the panel (explicit sides)
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
legend.position = "bottom", # "right" | "left" | "top" | "bottom"
legend.direction = "horizontal"
)
Suppress all legends
plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(x = "Years after Operation", y = "Freedom from Death (%)") +
hv_theme("poster")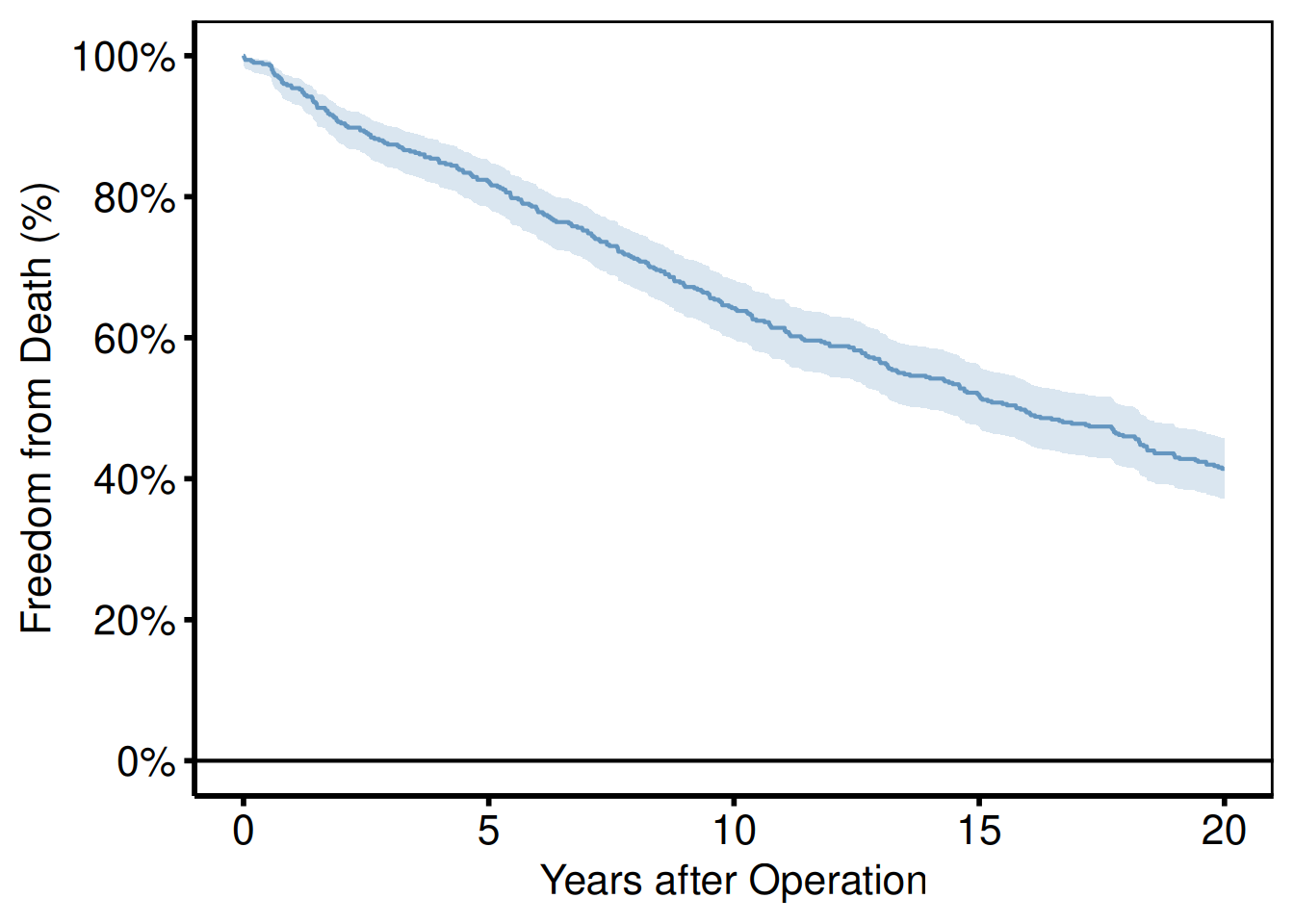
guide = "none" on every scale_*() call removes all legend entries. This is preferred over theme(legend.position = "none") when only some aesthetics have legends and others do not.
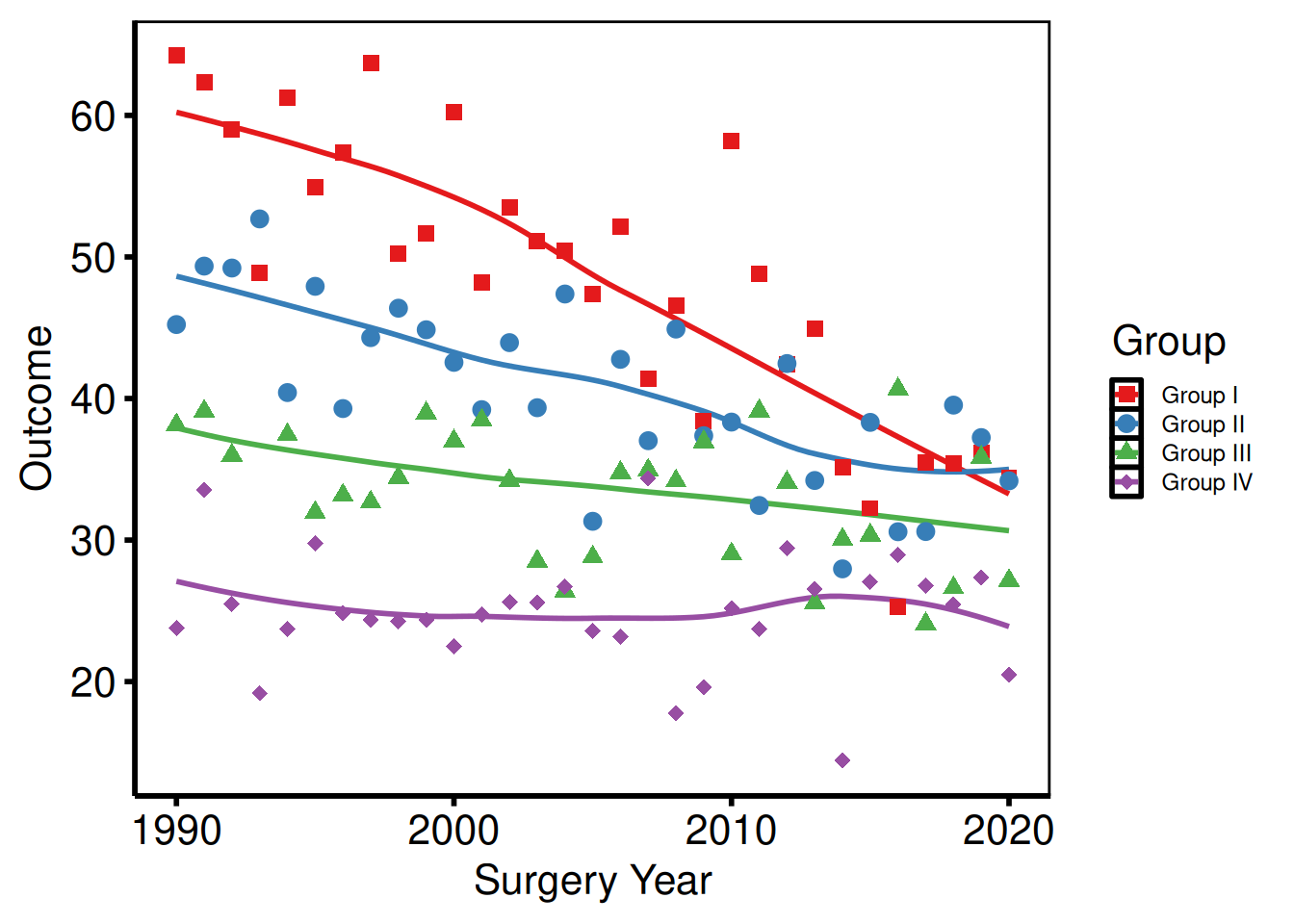
Legend text and key size
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
legend.text = element_text(size = 9),
legend.key.size = unit(0.4, "cm")
)
Theme Overrides
hv_theme() sets a complete non-data formatting baseline. Layer additional theme() calls after it to adjust individual elements without touching the rest.
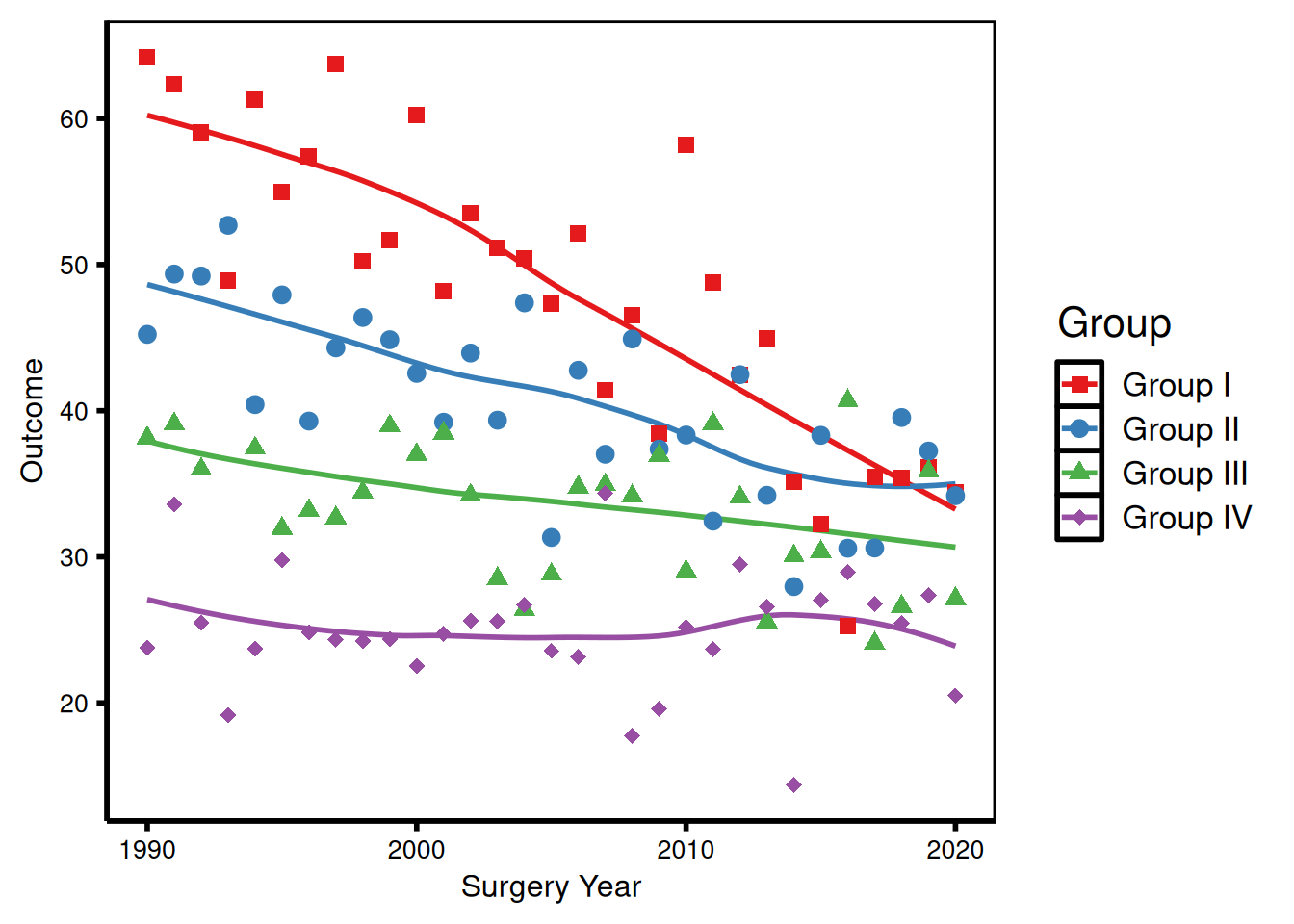
Axis text size
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
axis.text = element_text(size = 10), # tick labels
axis.title = element_text(size = 12) # axis titles
)
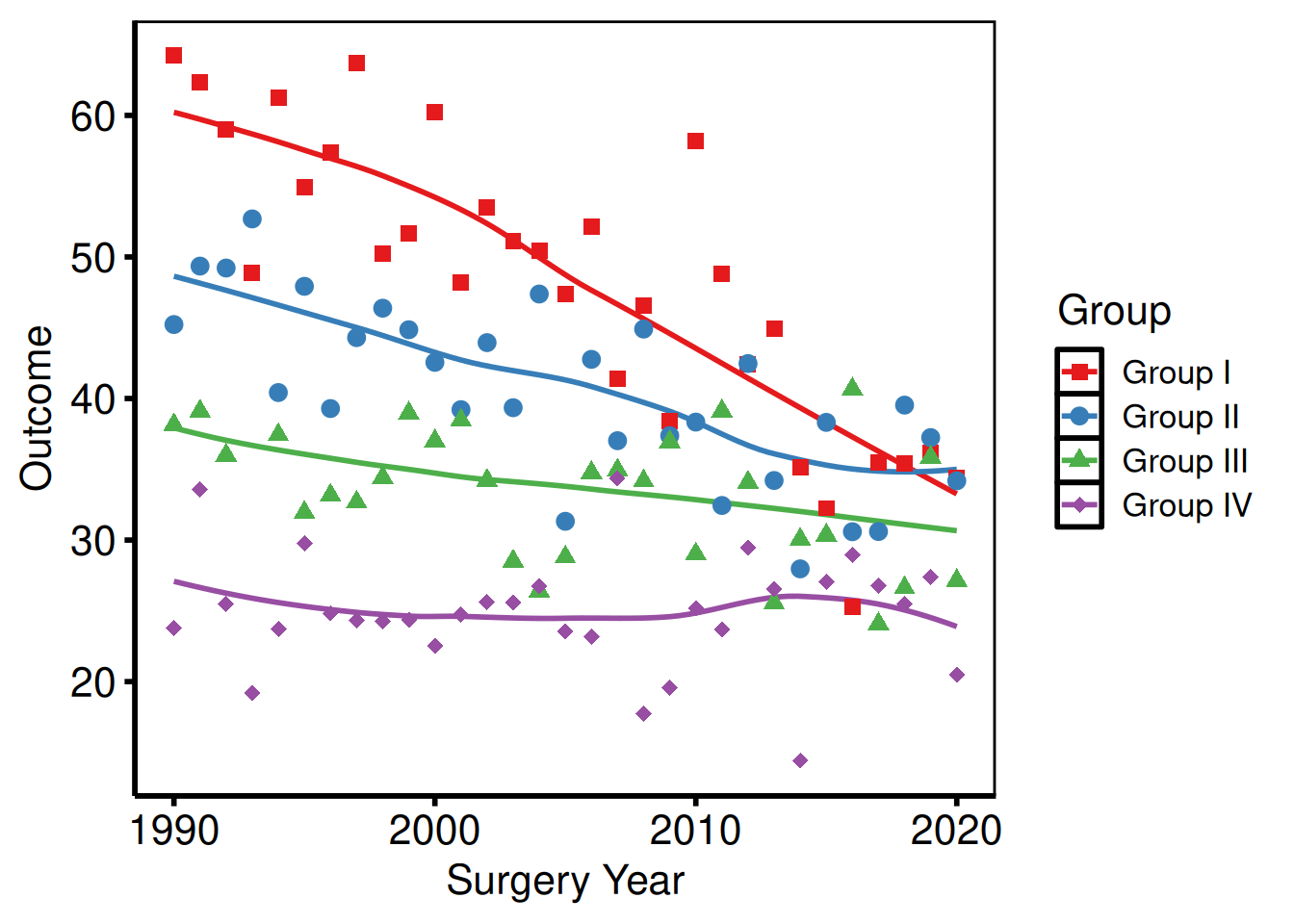
Removing minor grid lines
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
panel.grid.minor = element_blank()
)
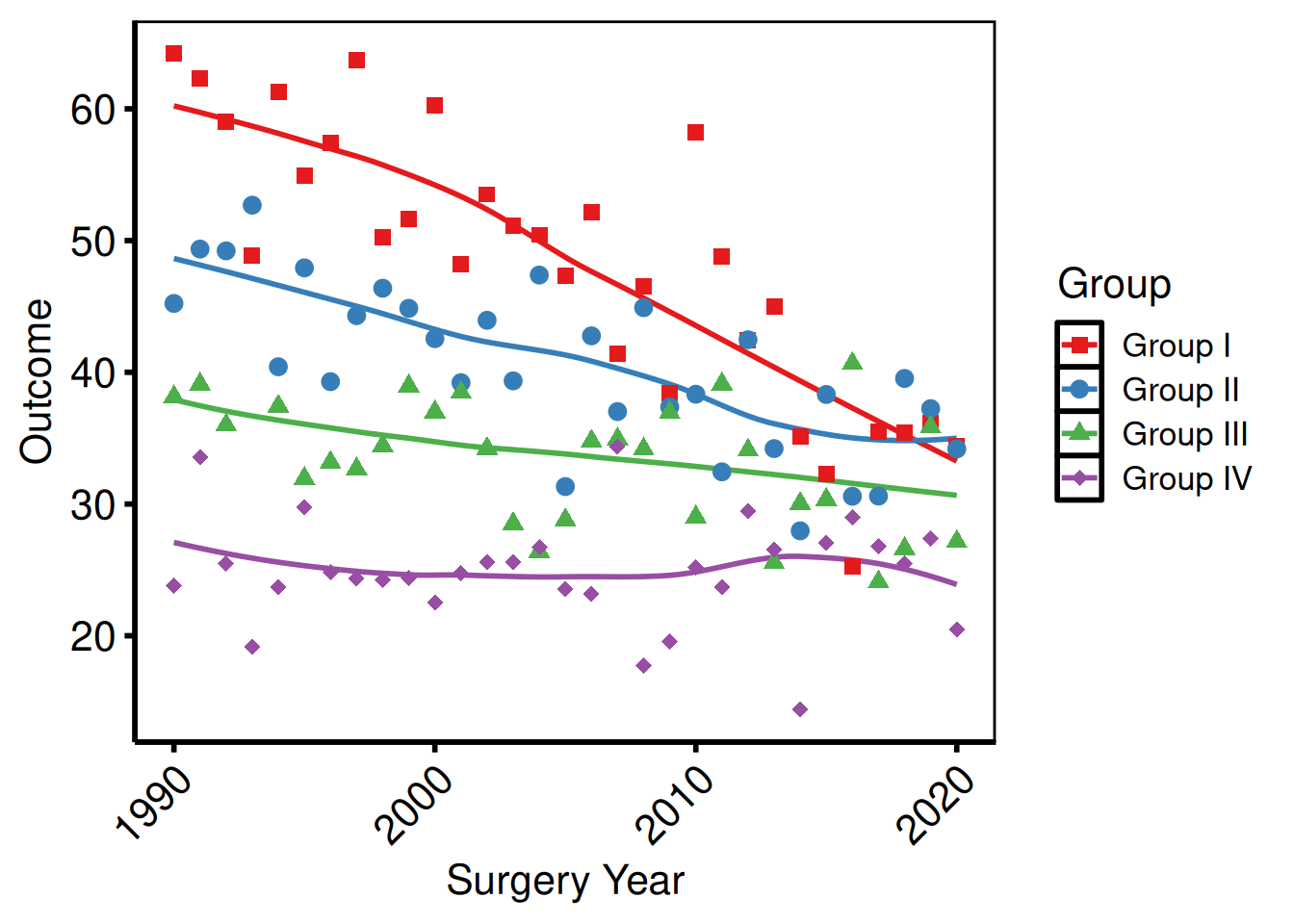
Rotating x-axis labels
Useful for time-point labels or long category names.
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1)
)
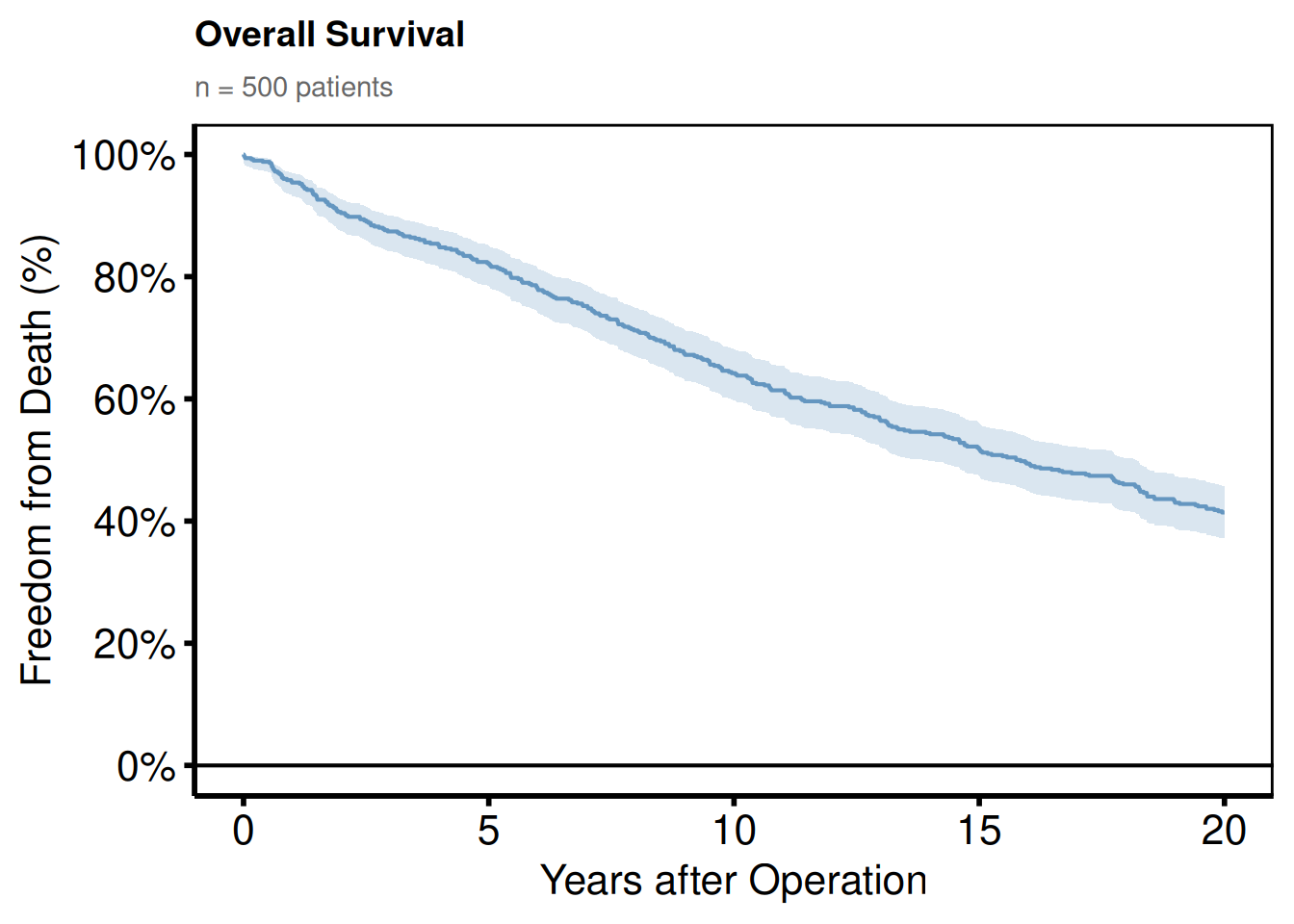
Adding a plot title and subtitle
Titles are stripped from the base themes (they are rarely used in journal figures), but can be added back:
plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(
title = "Overall Survival",
subtitle = paste0("n = ", nrow(dta_km), " patients"),
x = "Years after Operation",
y = "Freedom from Death (%)"
) +
hv_theme("poster") +
theme(
plot.title = element_text(size = 14, face = "bold", hjust = 0),
plot.subtitle = element_text(size = 11, colour = "grey40", hjust = 0)
)
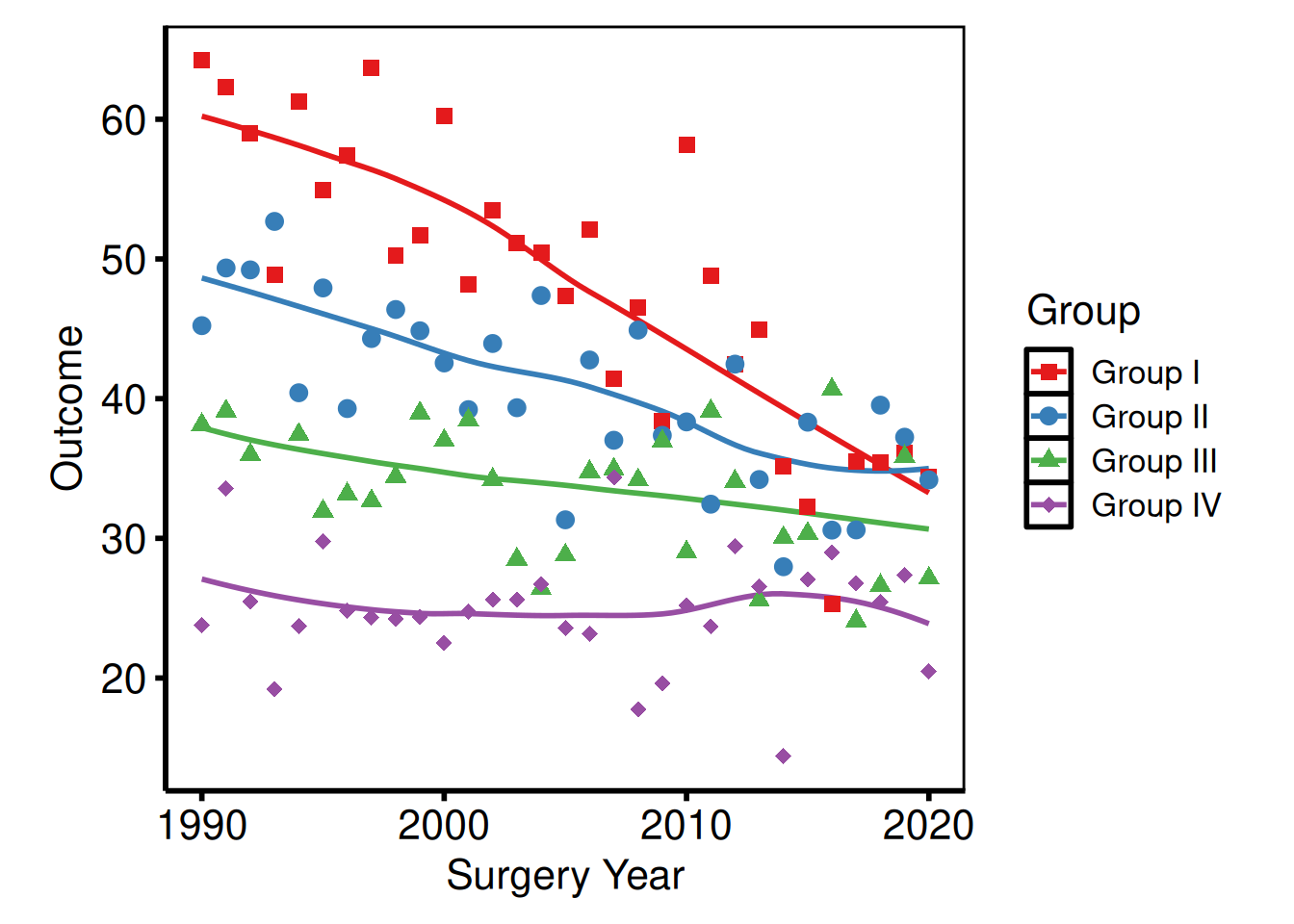
Expanding plot margins
Add breathing room around the panel, for example when a figure is placed directly on a poster without a surrounding text frame.
p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster") +
theme(
plot.margin = margin(t = 10, r = 20, b = 10, l = 20, unit = "pt")
)
Multi-panel Figures with patchwork
The patchwork package composes multiple ggplot objects into a single figure. Use it to place two related plots side by side, or to stack a main plot above a companion table or risk panel.
Side-by-side plots
library(patchwork)
p_ms <- p_base +
scale_colour_brewer(palette = "Set1", name = "Group") +
scale_shape_manual(
values = c("Group I" = 15, "Group II" = 19,
"Group III" = 17, "Group IV" = 18),
name = "Group"
) +
labs(x = "Surgery Year", y = "Outcome") +
hv_theme("poster")
p_km_ms <- plot(km) +
scale_color_manual(values = c(All = "steelblue"), guide = "none") +
scale_fill_manual(values = c(All = "steelblue"), guide = "none") +
scale_y_continuous(breaks = seq(0, 100, 20),
labels = function(x) paste0(x, "%")) +
scale_x_continuous(breaks = seq(0, 20, 5)) +
coord_cartesian(xlim = c(0, 20), ylim = c(0, 100)) +
labs(x = "Years after Operation", y = "Freedom from Death (%)") +
hv_theme("poster")
p_ms | p_km_ms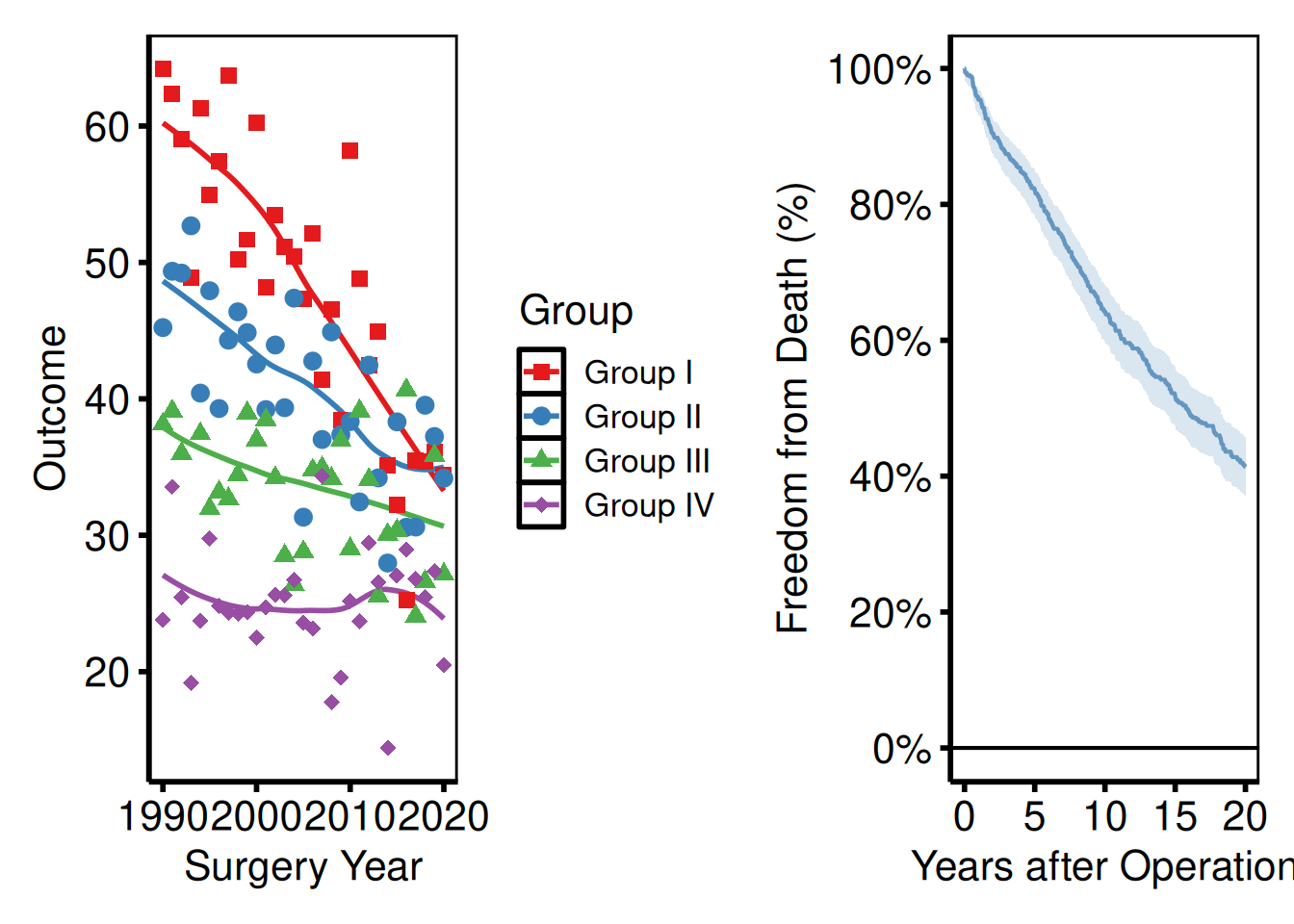
| places plots side by side; / stacks them vertically.
Controlling relative widths and heights
(p_ms | p_km_ms) +
plot_layout(widths = c(2, 1)) # left panel twice as wide as right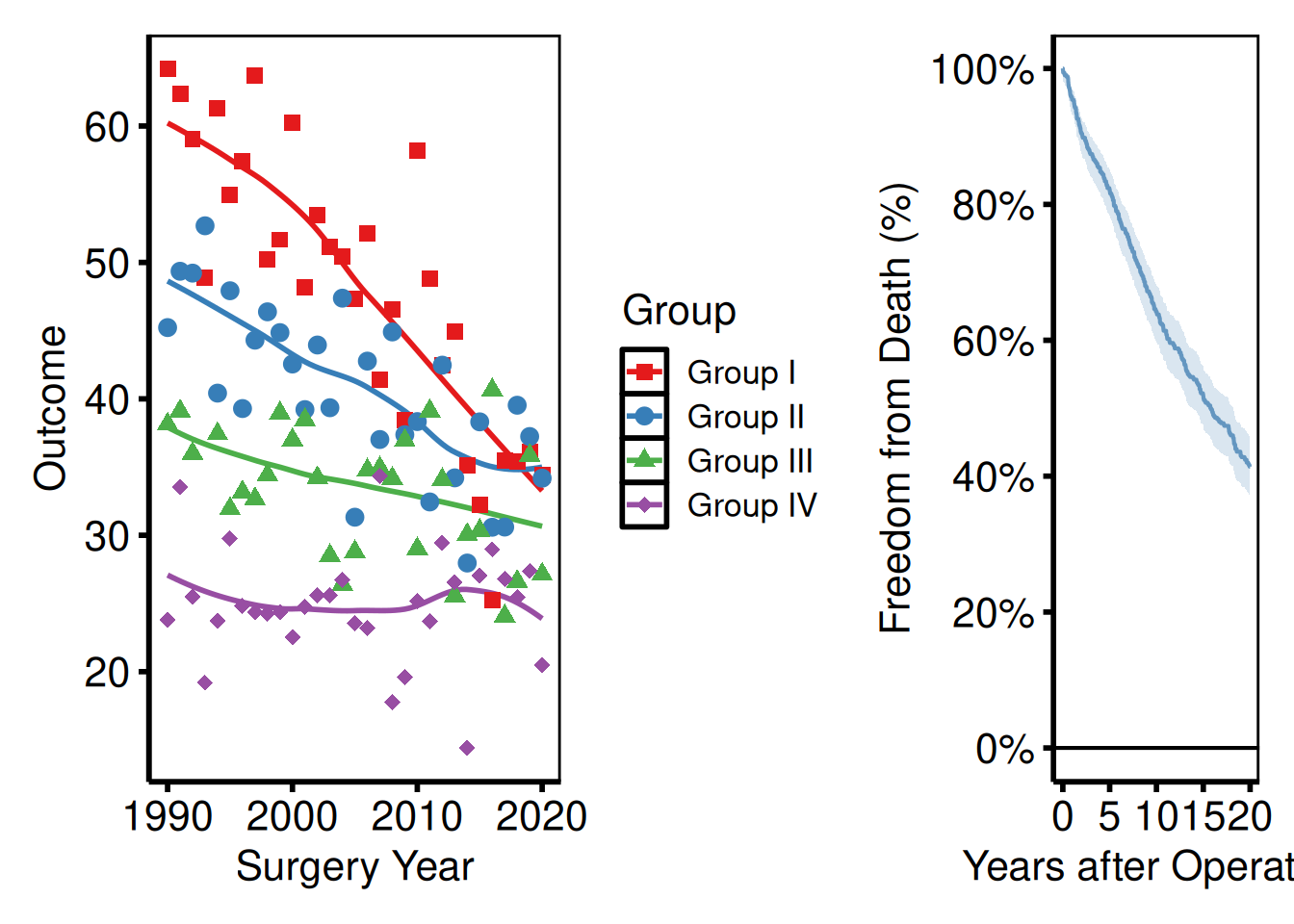
Stacking a plot above a risk table
A common pattern with survival curves is to pair the plot with a numbers-at-risk panel. hv_survival() stores the risk table as a data frame at km$tables$risk — columns strata, report_time, n.risk. Build a ggplot text panel from it, then stack with /.
risk_df <- km$tables$risk
rt_panel <- ggplot(risk_df,
aes(x = report_time, y = factor(strata),
label = n.risk)) +
geom_text(size = 3) +
scale_x_continuous(limits = c(0, 20), breaks = seq(0, 20, 5)) +
labs(x = "Years after Operation", y = NULL) +
hv_theme("poster") +
theme(
axis.line = element_blank(),
axis.ticks = element_blank()
)
p_km_ms / rt_panel +
plot_layout(heights = c(4, 1))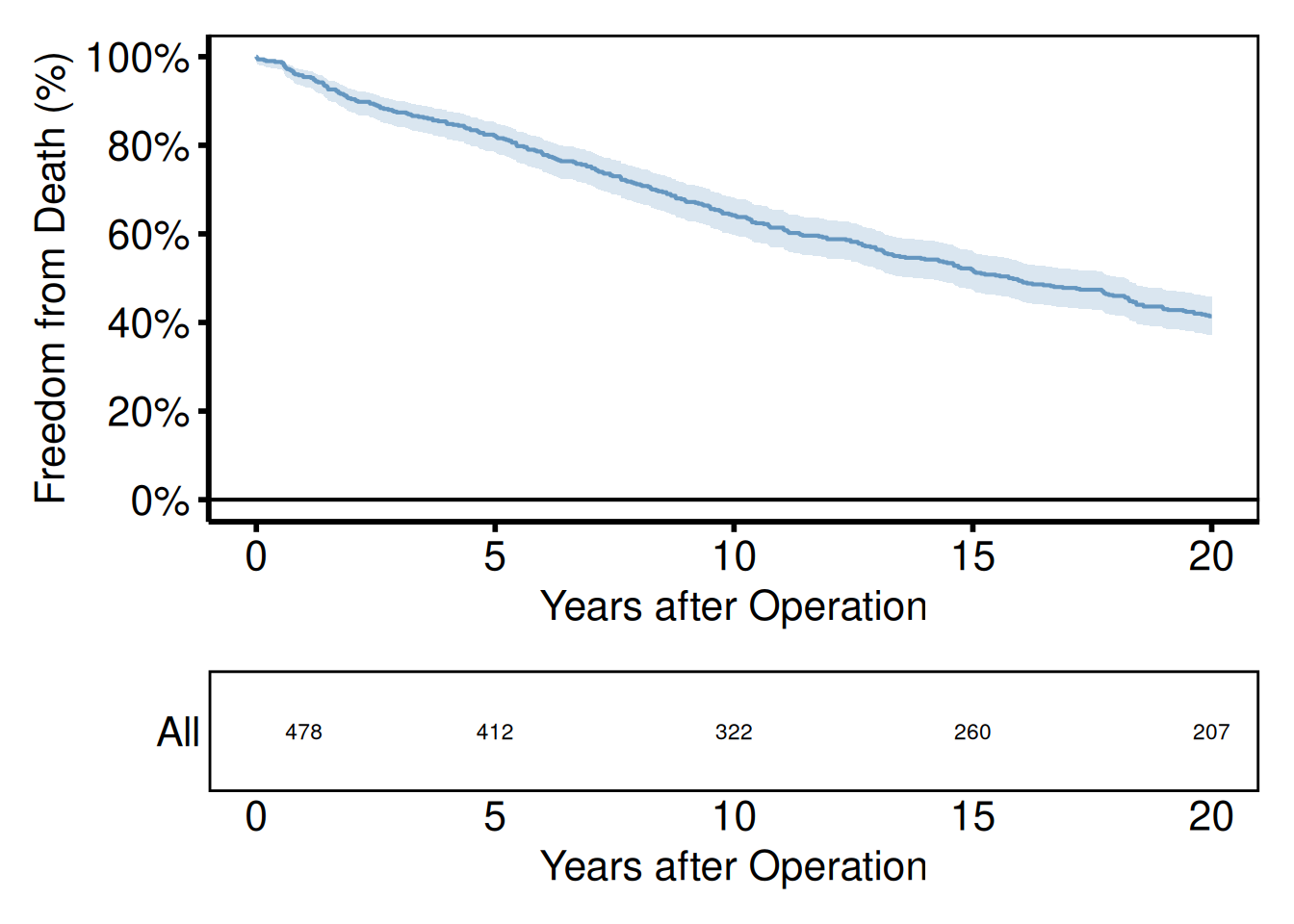
Shared axis labels and panel tags
plot_annotation() adds a shared title or tags (A, B, C…) across all panels.
(p_ms | p_km_ms) +
plot_annotation(
title = "Figure 1. Outcomes after cardiac surgery",
tag_levels = "A"
) &
theme(plot.tag = element_text(size = 12, face = "bold"))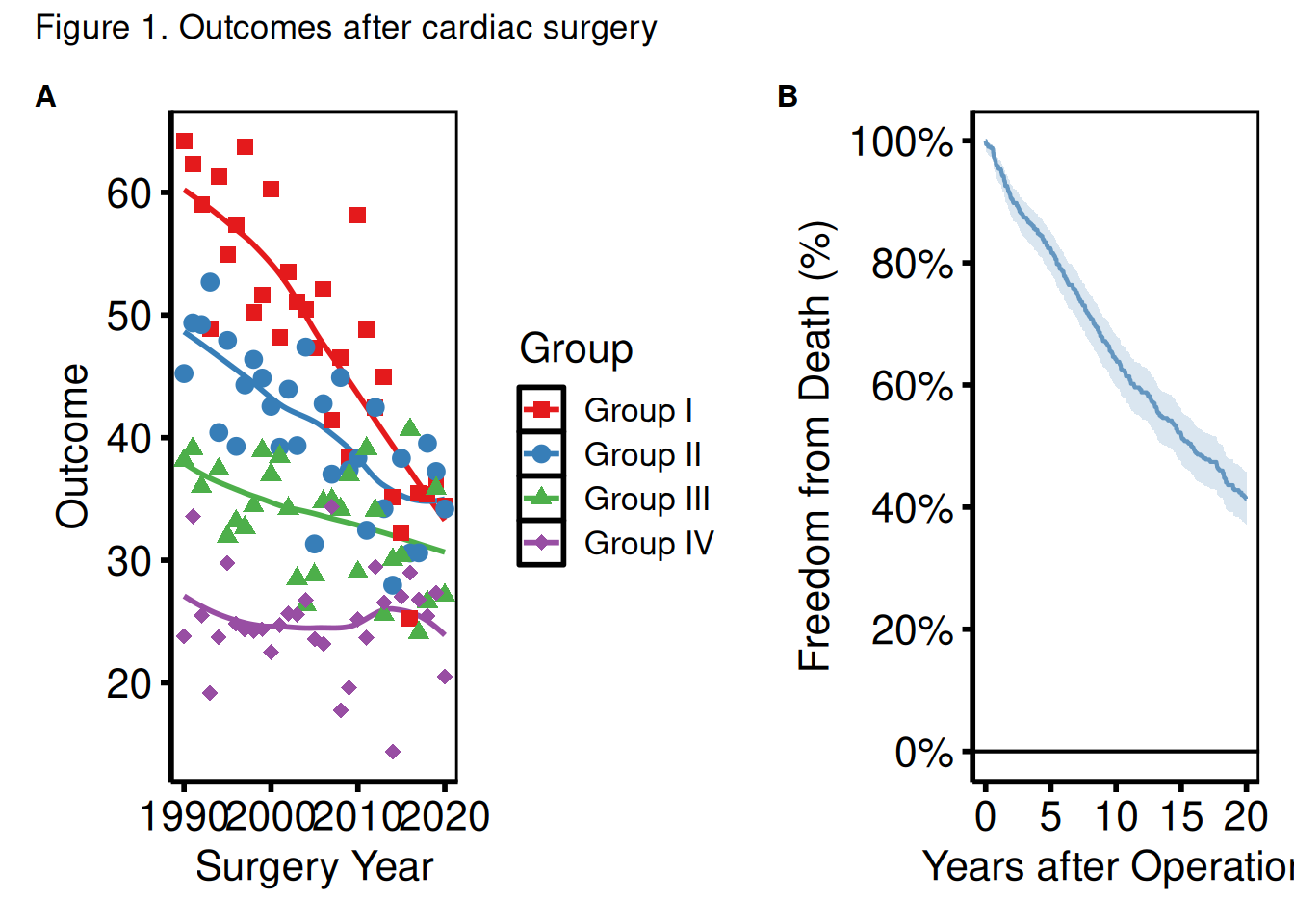
Saving a patchwork composite
Assign the composed object to a variable and pass it to ggsave(). For PowerPoint, save each panel individually with save_ppt() (patchwork composites are not editable DrawingML objects).
combined <- (p_ms | p_km_ms) +
plot_annotation(tag_levels = "A") &
theme(plot.tag = element_text(size = 12, face = "bold"))
ggsave(
filename = here::here("graphs", "fig1_combined.pdf"),
plot = combined,
width = 14,
height = 7
)