Fitting Hazard Models
From intercept-only to multiphase additive hazard
Source:vignettes/fitting-hazard-models.qmd
This vignette walks the core model-fitting workflow from the inside out: intercept-only fits to establish the baseline hazard shape, then covariates on top, then the multiphase decomposition, then multi-endpoint analyses on the same cohort. Every example uses a clinical dataset shipped with the package. If you haven’t seen the basics — what a parametric hazard model is, why we use the Surv(time, status) formula — start with vignette("getting-started") first; this vignette assumes that context.
The progression matters. Single-distribution intercept-only fits tell you whether the baseline hazard shape is monotone (Weibull territory) or has structure that demands a multiphase decomposition. Multivariable fits add covariate effects on top of a shape you already trust. Multiphase fits split the baseline shape into clinically interpretable phases. Multi-endpoint analyses reuse all the above for separate clinical outcomes — death, reoperation, infection — on the same patient cohort.
1 Intercept-only model: CABG survival (KU Leuven)
The cabgkul dataset contains 5,880 patients who underwent primary isolated coronary artery bypass grafting at KU Leuven between 1971 and 1987. With only two columns — follow-up time and death indicator — it is the simplest starting point.
Fit an intercept-only Weibull. With no covariates on the right-hand side of the formula the model estimates only the baseline hazard shape — the scale mu and exponent nu of a Weibull curve fit to all 5,880 patients pooled. This is the right starting point for any new dataset: before asking which covariates matter, ask whether a single monotone hazard even fits the population-level pattern.
The summary tells us where the optimizer landed; the picture tells us whether that landing point matches the data. Plot the fitted survival curve on a fine time grid and overlay the Kaplan-Meier step function from the raw cohort.
t_grid <- seq(0.01, max(cabgkul$int_dead) * 0.9, length.out = 200)
nd <- data.frame(time = t_grid)
surv <- predict(fit_kul, newdata = nd, type = "survival") * 100
km <- survfit(Surv(int_dead, dead) ~ 1, data = cabgkul)
km_df <- data.frame(time = km$time, survival = km$surv * 100)
ggplot() +
geom_step(data = km_df, aes(time, survival, colour = "Kaplan-Meier"),
linewidth = 0.5) +
geom_line(data = data.frame(time = t_grid, survival = surv),
aes(time, survival, colour = "Weibull"), linewidth = 1) +
scale_colour_manual(values = c("Weibull" = "#0072B2",
"Kaplan-Meier" = "#D55E00")) +
scale_y_continuous(limits = c(0, 100)) +
labs(x = "Months after CABG", y = "Freedom from death (%)",
colour = NULL) +
theme_minimal() +
theme(legend.position = "bottom")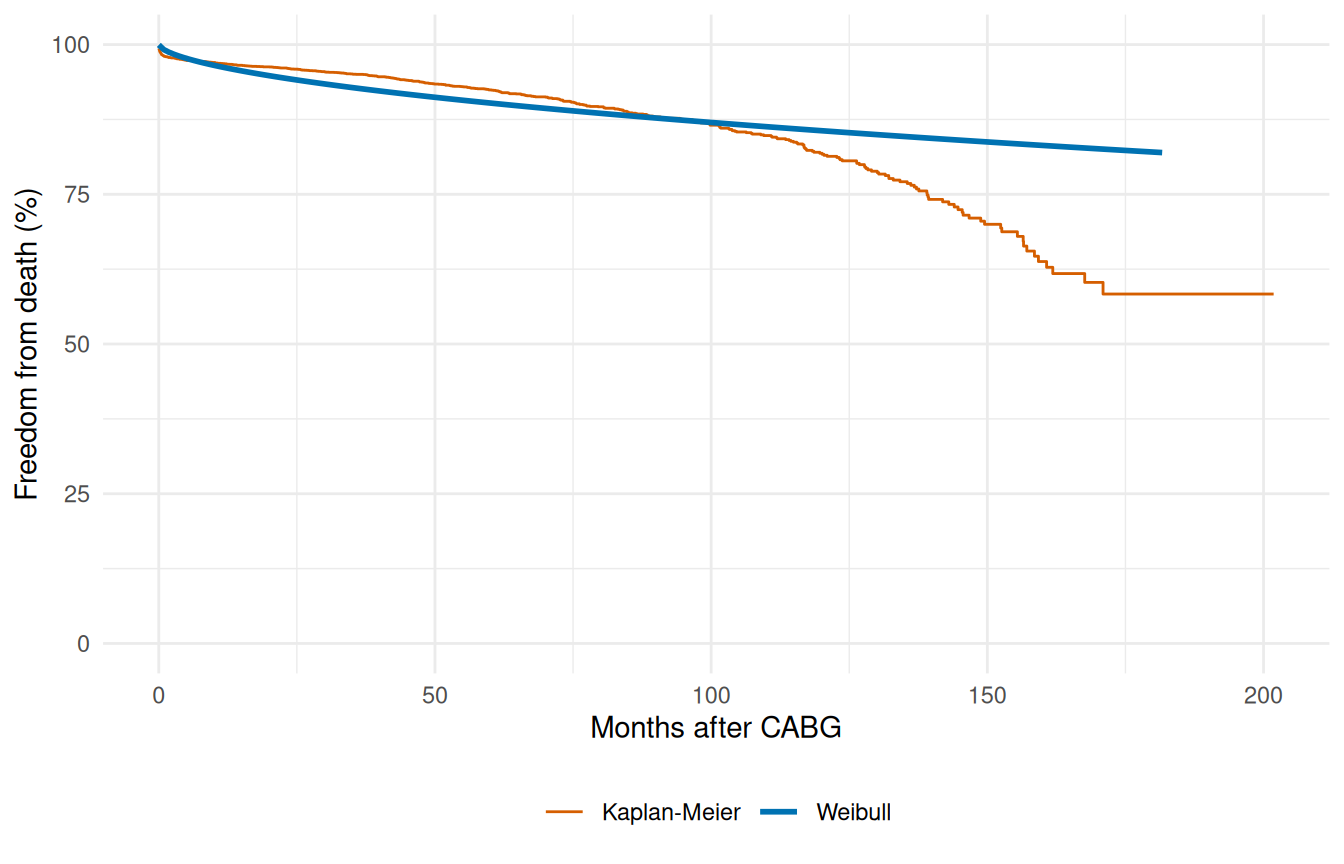
A single Weibull captures the broad trend but misses the distinct early operative risk and late attrition that the KM curve reveals. This motivates the multiphase approach below.
2 Multivariable model: AVC repair
The avc dataset has 310 patients who underwent atrioventricular canal repair, with 9 candidate covariates spanning patient demographics (age, NYHA status), anatomical features (malalignment, orifice morphology), intra-operative grading (inc_surg = surgical grade of AV valve incompetence), and post-operative complications (com_iv = grade IV complications). We drop incomplete rows so the design matrix is rectangular, then look at the column types and ranges.
data(avc)
avc <- na.omit(avc)
str(avc)
#> 'data.frame': 305 obs. of 11 variables:
#> $ study : chr "001C" "002C" "004C" "005C" ...
#> $ status : int 3 3 1 2 2 3 1 1 3 3 ...
#> $ inc_surg: int 4 3 2 3 1 2 3 2 3 3 ...
#> $ opmos : num 9.46 34.07 51.58 55 60.65 ...
#> $ age : num 69.2 53.7 286.1 154.6 48.4 ...
#> $ mal : int 0 0 0 1 0 0 0 0 0 0 ...
#> $ com_iv : int 1 1 1 1 1 1 1 1 1 1 ...
#> $ orifice : int 0 0 0 0 0 0 0 0 0 0 ...
#> $ dead : int 1 1 0 0 0 0 0 0 0 1 ...
#> $ int_dead: num 0.0534 0.3778 91.5337 111.608 106.8112 ...
#> $ op_age : num 654 1828 14759 8505 2933 ...
#> - attr(*, "na.action")= 'omit' Named int [1:5] 12 90 138 144 146
#> ..- attr(*, "names")= chr [1:5] "12" "90" "138" "144" ...Now we put covariates on the right-hand side of the formula and refit. The theta vector grows: two Weibull shape parameters (mu, nu) plus six covariate coefficients (beta1..beta6), each starting at zero. The optimizer estimates a log-hazard-ratio for every covariate jointly with the Weibull shape — so the shape and the covariate effects are identified from the same likelihood, not sequentially.
fit_avc <- hazard(
Surv(int_dead, dead) ~ age + status + mal + com_iv + inc_surg + orifice,
data = avc,
dist = "weibull",
theta = c(mu = 0.20, nu = 1.0, rep(0, 6)),
fit = TRUE,
control = list(maxit = 500)
)
fit_avc
#> hazard object
#> observations: 305
#> predictors: 6
#> dist: weibull
#> engine: native-r-m2
#> log-lik: -197.159
#> converged: TRUEEach coefficient is a log-hazard-ratio: positive means higher risk, negative means lower, zero means no effect. The large positive coefficients on mal (anatomical malalignment) and com_iv (grade IV post-operative complications) flag these as the dominant risk markers in this cohort. The standard errors and Wald z-statistics in the summary tell you which effects are well identified and which are noise — a coefficient with a z-statistic near zero contributes essentially nothing the data can defend.
3 Multiphase model: additive hazard decomposition
The single-Weibull fits above gave us a curve that’s mediocre everywhere instead of right anywhere. That’s a structural limitation of a monotone parametric shape, not something more iterations will fix. The Blackstone–Naftel–Turner framework’s key idea is to split the hazard into a sum of phase-specific contributions, each with its own temporal shape and its own scale:
Each is a phase-specific unit-scaled curve (early-peaking saturating, flat constant, late-rising polynomial) and each is the phase-specific scale, possibly modulated by covariates. The phases overlap and add — no switching, no thresholds — so the total instantaneous hazard at any is the sum of the per-phase rates. See vignette("getting-started") for the longer-form motivation; what follows here is the practical workflow for fitting one.
For AVC we’ll use two phases — an early phase to absorb the operative-window mortality, and a constant phase for the background rate. AVC patients don’t have a clear late-deterioration regime over this follow-up window, so a third (g3) phase would be unidentified. We fix the shape parameters and estimate only the scales, matching the workflow you’d run against a SAS HAZARD reference fit.
fit_mp <- hazard(
Surv(int_dead, dead) ~ 1,
data = avc,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 1, m = 1,
fixed = "shapes"),
constant = hzr_phase("constant")
),
fit = TRUE,
control = list(n_starts = 5, maxit = 1000)
)
summary(fit_mp)
#> Multiphase hazard model (2 phases)
#> observations: 305
#> predictors: 0
#> dist: multiphase
#> phase 1: early - cdf (early risk)
#> phase 2: constant - constant (flat rate)
#> engine: native-r-m2
#> converged: TRUE
#> log-lik: -228.029
#> evaluations: fn=32, gr=10
#>
#> Coefficients (internal scale):
#>
#> Phase: early (cdf)
#> estimate std_error z_stat p_value
#> log_mu -1.4132735 0.04410012 -32.04693 2.422767e-225
#> log_t_half -0.6931472 NA NA NA
#> nu 1.0000000 NA NA NA
#> m 1.0000000 NA NA NA
#>
#> Phase: constant (constant)
#> estimate std_error z_stat p_value
#> log_mu -7.609476 NA NA NAThe diagnostic that matters is whether the multiphase fit actually out-performs the single Weibull against the data. Plot both parametric curves against the same Kaplan-Meier reference so we can see, by eye, where each model is honest and where each is reaching.
t_grid <- seq(0.01, max(avc$int_dead) * 0.95, length.out = 200)
nd <- data.frame(time = t_grid)
km_avc <- survfit(Surv(int_dead, dead) ~ 1, data = avc)
km_df <- data.frame(time = km_avc$time, survival = km_avc$surv * 100)
fit_wb <- hazard(
Surv(int_dead, dead) ~ 1, data = avc, dist = "weibull",
theta = c(mu = 0.20, nu = 1.0), fit = TRUE
)
surv_wb <- predict(fit_wb, newdata = nd, type = "survival") * 100
surv_mp <- predict(fit_mp, newdata = nd, type = "survival") * 100
plot_df <- rbind(
data.frame(time = t_grid, survival = surv_wb, Model = "Single Weibull"),
data.frame(time = t_grid, survival = surv_mp, Model = "Multiphase (2-phase)")
)
ggplot() +
geom_step(data = km_df, aes(time, survival), colour = "grey50",
linewidth = 0.5) +
geom_line(data = plot_df, aes(time, survival, colour = Model),
linewidth = 1) +
scale_colour_manual(values = c("Single Weibull" = "#E69F00",
"Multiphase (2-phase)" = "#0072B2")) +
scale_y_continuous(limits = c(0, 100)) +
annotate("text", x = max(t_grid) * 0.6, y = 95, label = "KM (grey)",
size = 3, colour = "grey50") +
labs(x = "Months after AVC repair", y = "Freedom from death (%)",
colour = NULL) +
theme_minimal() +
theme(legend.position = "bottom")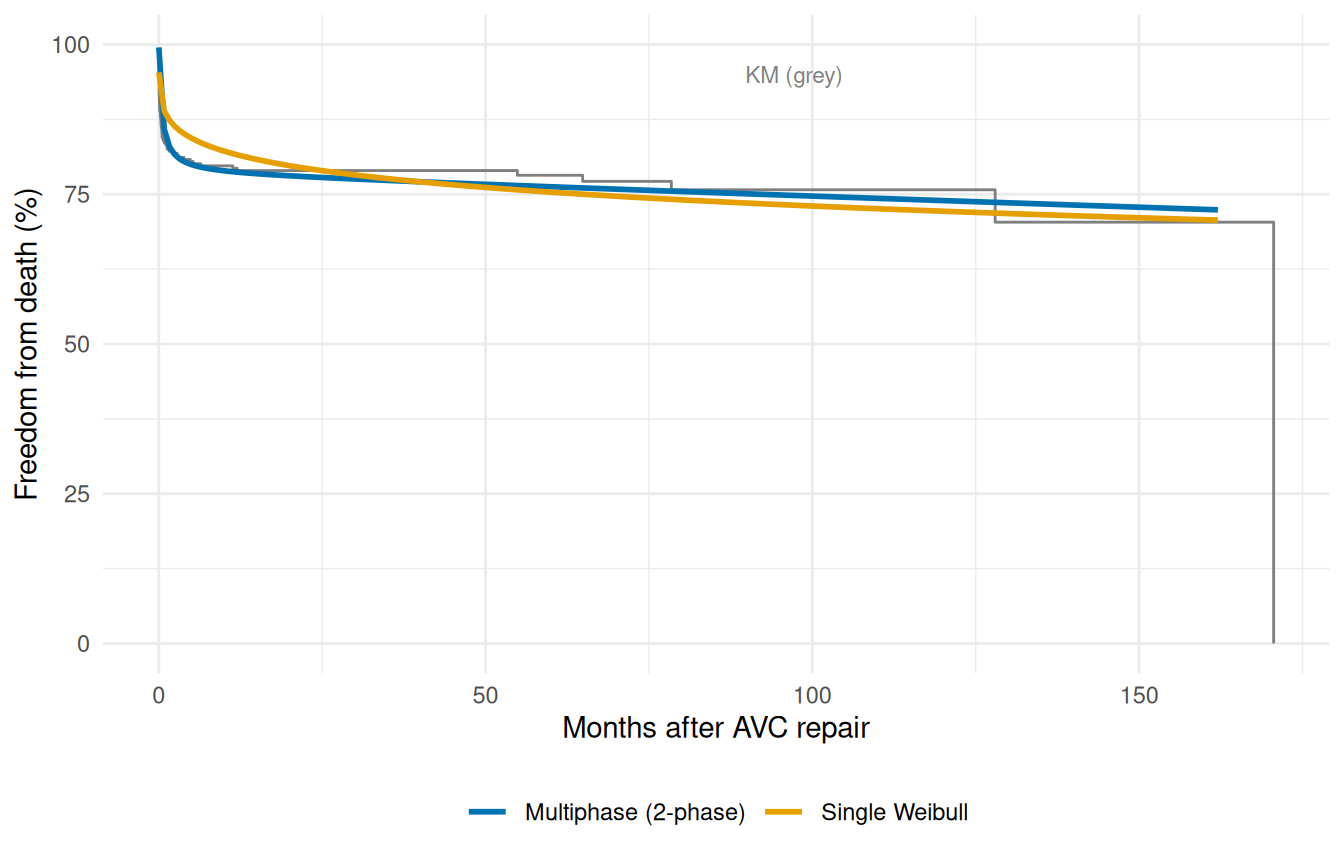
The multiphase model tracks the KM curve much more closely than the single Weibull, especially across the steep early-mortality window — which is exactly where the single Weibull was forced to compromise. The constant phase then carries the slow post-recovery attrition. The point isn’t that multiphase always wins; it’s that when the data has phase structure, fitting that structure explicitly is strictly more honest than averaging it away into one monotone curve.
4 Multi-endpoint models: heart valve replacement
The valves dataset (1,533 patients) has multiple time-to-event endpoints — death, prosthetic valve endocarditis (PVE), and reoperation — each with its own follow-up time and event indicator. The same hazard() call fits each endpoint independently:
Start with the death endpoint. We use age at operation, NYHA class, and mechanical-valve indicator as covariates — the clinically canonical set for survival after valve replacement.
data(valves)
valves <- na.omit(valves)
fit_death <- hazard(
Surv(int_dead, dead) ~ age_cop + nyha + mechvalv,
data = valves,
dist = "weibull",
theta = c(mu = 0.10, nu = 1.0, rep(0, 3)),
fit = TRUE,
control = list(maxit = 500)
)
fit_death
#> hazard object
#> observations: 1523
#> predictors: 3
#> dist: weibull
#> engine: native-r-m2
#> log-lik: -1820.55
#> converged: TRUESwitch endpoints. Same data, same package, but now we model time to prosthetic valve endocarditis instead of death. The covariate list shifts to match the clinical question: nve (native-valve endocarditis history) replaces nyha because functional class is less informative for infection risk than prior endocarditis exposure. The fit returns its own MLE, coefficients, and standard errors completely independent of the death model.
fit_pve <- hazard(
Surv(int_pve, pve) ~ age_cop + nve + mechvalv,
data = valves,
dist = "weibull",
theta = c(mu = 0.02, nu = 1.0, rep(0, 3)),
fit = TRUE,
control = list(maxit = 500)
)
fit_pve
#> hazard object
#> observations: 1523
#> predictors: 3
#> dist: weibull
#> engine: native-r-m2
#> log-lik: -391.125
#> converged: TRUEEach endpoint gets its own model with its own covariates, but the hazard model structure — temporal shape plus covariate effects — stays the same whatever the clinical endpoint. Repeating the workflow for a third endpoint (reoperation, for example) is mechanical: swap the Surv(...) columns, swap the covariates, refit. The advantage over running three separate analyses in different tools is that the predictions, diagnostics, and uncertainty quantification all come from the same package — there’s no risk of subtle differences in censoring handling or estimator choice between endpoints.
5 Phase types reference
You’ve now seen each phase type in use: a "cdf" early phase for AVC operative mortality, a "constant" phase for AVC background rate, and the implicit single shape of every Weibull fit. The package supports three phase types in total, summarized here for quick reference:
| Type | Description | Typical use |
|---|---|---|
"cdf" |
Sigmoidal CDF shape (parameterized by t_half, nu, m) |
Early or late phases with transient risk |
"constant" |
Flat hazard (no temporal shape parameters) | Ongoing background risk |
"g3" |
Late-phase G3 parameterization (4 parameters: tau, gamma, alpha, eta) |
Late-rising risk matching C/SAS G3 output |
The "cdf" type covers the widest range of shapes: setting t_half small (e.g., 0.5) creates an early-peaking phase; setting it large (e.g., 10) creates a late-rising phase. The "constant" phase needs no shape parameters. The "g3" shape is the explicit late-rising parameterization that matches the SAS HAZARD “late” library; use it when you need parity against a C/SAS reference fit, or when the late rise has a clear lag-then-accelerate pattern that a delayed "cdf" doesn’t capture cleanly. See vignette("mf-mathematical-foundations") for the full mathematical treatment of each, including the parameter identifiability constraints.