library(TemporalHazard)
if (has_ggplot2) library(ggplot2)
data(avc)
avc <- na.omit(avc)
str(avc)
#> 'data.frame': 305 obs. of 11 variables:
#> $ study : chr "001C" "002C" "004C" "005C" ...
#> $ status : int 3 3 1 2 2 3 1 1 3 3 ...
#> $ inc_surg: int 4 3 2 3 1 2 3 2 3 3 ...
#> $ opmos : num 9.46 34.07 51.58 55 60.65 ...
#> $ age : num 69.2 53.7 286.1 154.6 48.4 ...
#> $ mal : int 0 0 0 1 0 0 0 0 0 0 ...
#> $ com_iv : int 1 1 1 1 1 1 1 1 1 1 ...
#> $ orifice : int 0 0 0 0 0 0 0 0 0 0 ...
#> $ dead : int 1 1 0 0 0 0 0 0 0 1 ...
#> $ int_dead: num 0.0534 0.3778 91.5337 111.608 106.8112 ...
#> $ op_age : num 654 1828 14759 8505 2933 ...
#> - attr(*, "na.action")= 'omit' Named int [1:5] 12 90 138 144 146
#> ..- attr(*, "names")= chr [1:5] "12" "90" "138" "144" ...Complete Clinical Analysis Walkthrough
From Kaplan-Meier baseline to validated multivariable model
2026-06-02
Source:vignettes/clinical-analysis-walkthrough.qmd
The previous vignettes covered each piece of the analytical workflow in isolation — how to fit a model, how to predict from it, how to validate it. This vignette runs all of those pieces together on a single dataset, in the order you’d actually run them in a real clinical analysis. The point isn’t to introduce new functions; it’s to show how the pieces chain into a disciplined sequence that turns raw covariates into a fitted, validated, defensible hazard model.
The sequence mirrors the one in the original SAS HAZARD system, which codified what cardiothoracic-surgery biostatisticians had been doing informally for decades:
- Nonparametric baseline — Kaplan-Meier life table to see what the data is saying before any model intervenes.
- Shape fitting — start with a single-distribution Weibull, build up to a multiphase decomposition when the KM curve demands it.
- Variable screening — univariable association tests and calibration plots to decide which covariates enter the model and in what functional form.
- Multivariable model — covariates layered onto a hazard shape you already trust.
- Prediction — patient-specific risk profiles for clinical reporting and decision support.
- Validation — decile-of-risk calibration to verify the model is honest across the risk spectrum, not just on average.
This corresponds to the SAS programs ac.* → hz.* → lg.* → hm.* → hp.* → hs.*. If you’re familiar with the SAS workflow, the section structure should feel familiar; if not, treat this as the canonical analytical sequence to follow on any new dataset.
We use the AVC dataset (310 patients, atrioventricular canal repair) which has rich covariates and two identifiable hazard phases — fewer phases than the 3-phase CABG example you’ve seen in other vignettes, but with the covariate complexity needed to exercise the screening, multivariable-fit, and validation steps.
1 Data preparation
Load the package and the AVC dataset, drop incomplete rows so the design matrix is rectangular for the multivariable fits to come, and inspect the resulting column types and ranges. The na.omit() step is conservative — losing rows is a real cost — but for a walkthrough we want every later fit to use the same patient set so the comparisons are apples-to-apples.
The AVC dataset contains 305 patients after removing incomplete cases. Key covariates include age at repair (age, in months), NYHA functional class (status), presence of malalignment (mal), and interventricular communication (com_iv).
2 Step 1: Nonparametric baseline
Before fitting any parametric model, establish the empirical survival curve using the Kaplan-Meier estimator. This is the benchmark against which all parametric fits will be compared.
hzr_kaplan() wraps survival::survfit() and adds logit-transformed exact confidence limits that respect the [0, 1] boundary — more accurate in the tails than the default Greenwood intervals. This matches the SAS kaplan.sas macro output structure.
km <- hzr_kaplan(time = avc$int_dead, status = avc$dead)
head(km)
#> Kaplan-Meier estimate with logit confidence limits
#> Events: 18 | Time points: 6
#> Survival range: 0.941 to 0.9836
#> RMST at last event: 0.0472
#>
#> time n_risk n_event n_censor survival std_err cl_lower cl_upper cumhaz
#> 0.001368954 305 5 0 0.9836 0.0073 0.9612 0.9932 0.0165
#> 0.002737907 300 2 0 0.9770 0.0086 0.9527 0.9890 0.0232
#> 0.008213721 298 2 0 0.9705 0.0097 0.9443 0.9846 0.0300
#> 0.016427440 296 5 0 0.9541 0.0120 0.9240 0.9726 0.0470
#> 0.032854880 291 3 0 0.9443 0.0131 0.9122 0.9651 0.0574
#> 0.049282330 288 1 0 0.9410 0.0135 0.9083 0.9625 0.0608
#> hazard density life
#> 12.0744 11.9752 0.0014
#> 4.8862 4.7901 0.0027
#> 1.2298 1.1975 0.0081
#> 2.0741 1.9959 0.0160
#> 0.6308 0.5988 0.0317
#> 0.2117 0.1996 0.0472The returned data frame is the life table: one row per event time with n_risk, n_event, survival, logit-transformed 95% CLs (cl_lower / cl_upper), and a KM-based cumulative hazard (-log(survival) — use hzr_nelson() if you need the Nelson-Aalen estimator instead). Plotting just requires the survival column:
km_df <- data.frame(
time = km$time,
survival = km$survival * 100,
lower = km$cl_lower * 100,
upper = km$cl_upper * 100
)
ggplot(km_df, aes(time, survival)) +
geom_step(colour = "#D55E00", linewidth = 0.8) +
geom_ribbon(aes(ymin = lower, ymax = upper),
stat = "identity", alpha = 0.15, fill = "#D55E00") +
labs(x = "Months after AVC repair", y = "Survival (%)") +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal()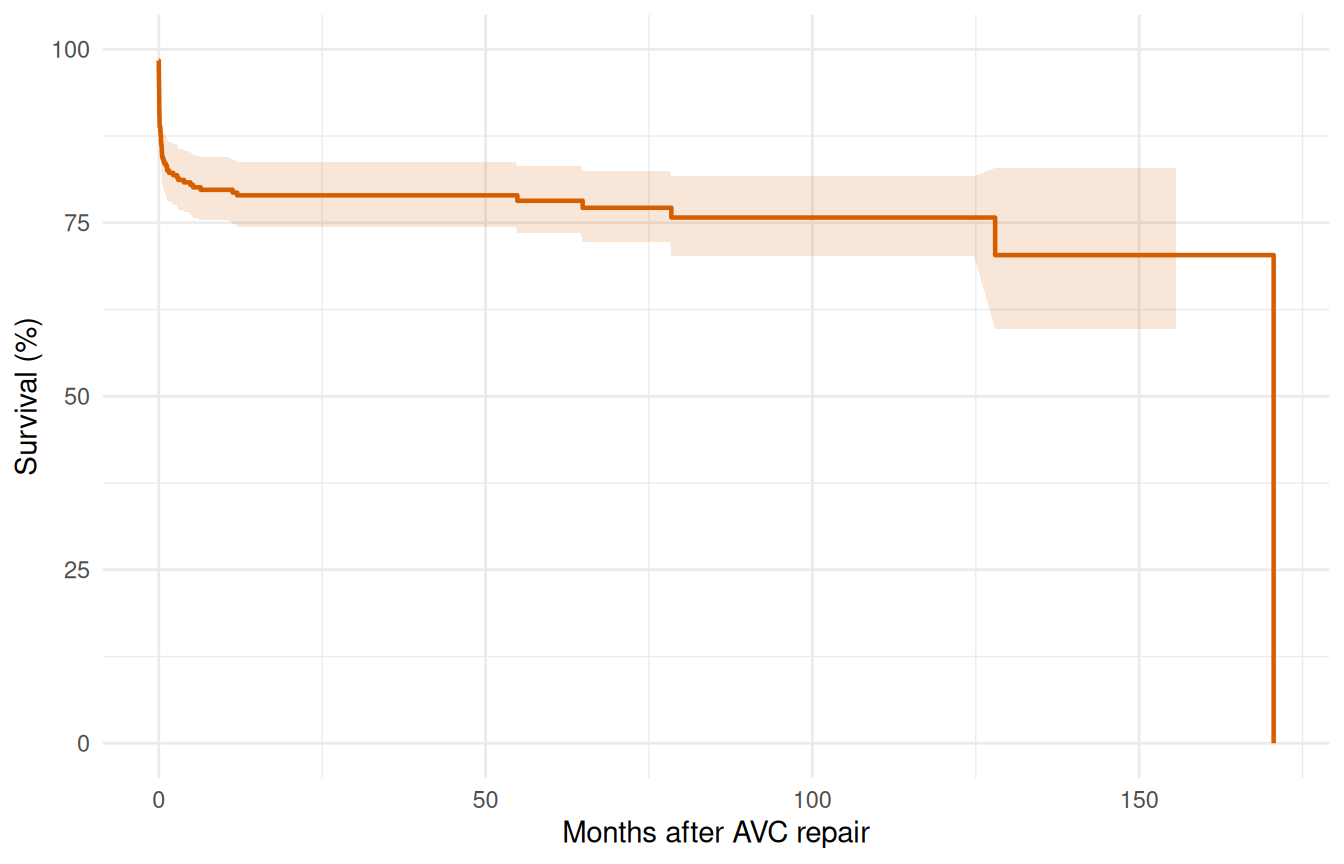
The KM curve shows a sharp early drop (operative mortality) followed by a roughly constant attrition rate. This two-phase pattern suggests an early CDF phase plus a constant phase — no obvious late rising hazard.
3 Step 2: Shape fitting — simple to complex
3.1 2a. Single-phase Weibull
The discipline here is to start with the simplest parametric model and earn any added complexity. A single-distribution Weibull intercept-only fit gives us a smooth two-parameter curve to compare against the KM baseline. If the Weibull tracks the KM step function reasonably well, we may be done; if it can’t, the gap tells us exactly where a richer model needs to go.
fit_weib <- hazard(
survival::Surv(int_dead, dead) ~ 1,
data = avc,
dist = "weibull",
theta = c(mu = 0.05, nu = 0.5),
fit = TRUE
)
summary(fit_weib)
#> hazard model summary
#> observations: 305
#> predictors: 0
#> dist: weibull
#> engine: native-r-m2
#> converged: TRUE
#> log-lik: -234.775
#> evaluations: fn=34, gr=9
#>
#> Coefficients:
#> estimate std_error z_stat p_value
#> mu 3.498711e-05 0.000034554 1.012534 3.112826e-01
#> nu 2.043444e-01 0.023324005 8.761118 1.933229e-18The Weibull forces a monotone hazard shape. Let’s overlay it on the Kaplan-Meier to see where it fits well and where it doesn’t.
t_grid <- seq(0.01, max(avc$int_dead) * 0.9, length.out = 200)
surv_weib <- predict(fit_weib,
newdata = data.frame(time = t_grid),
type = "survival") * 100
ggplot() +
geom_step(data = km_df, aes(time, survival),
colour = "#D55E00", linewidth = 0.6) +
geom_line(data = data.frame(time = t_grid, survival = surv_weib),
aes(time, survival), colour = "#0072B2", linewidth = 0.8) +
labs(x = "Months after AVC repair", y = "Survival (%)") +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal()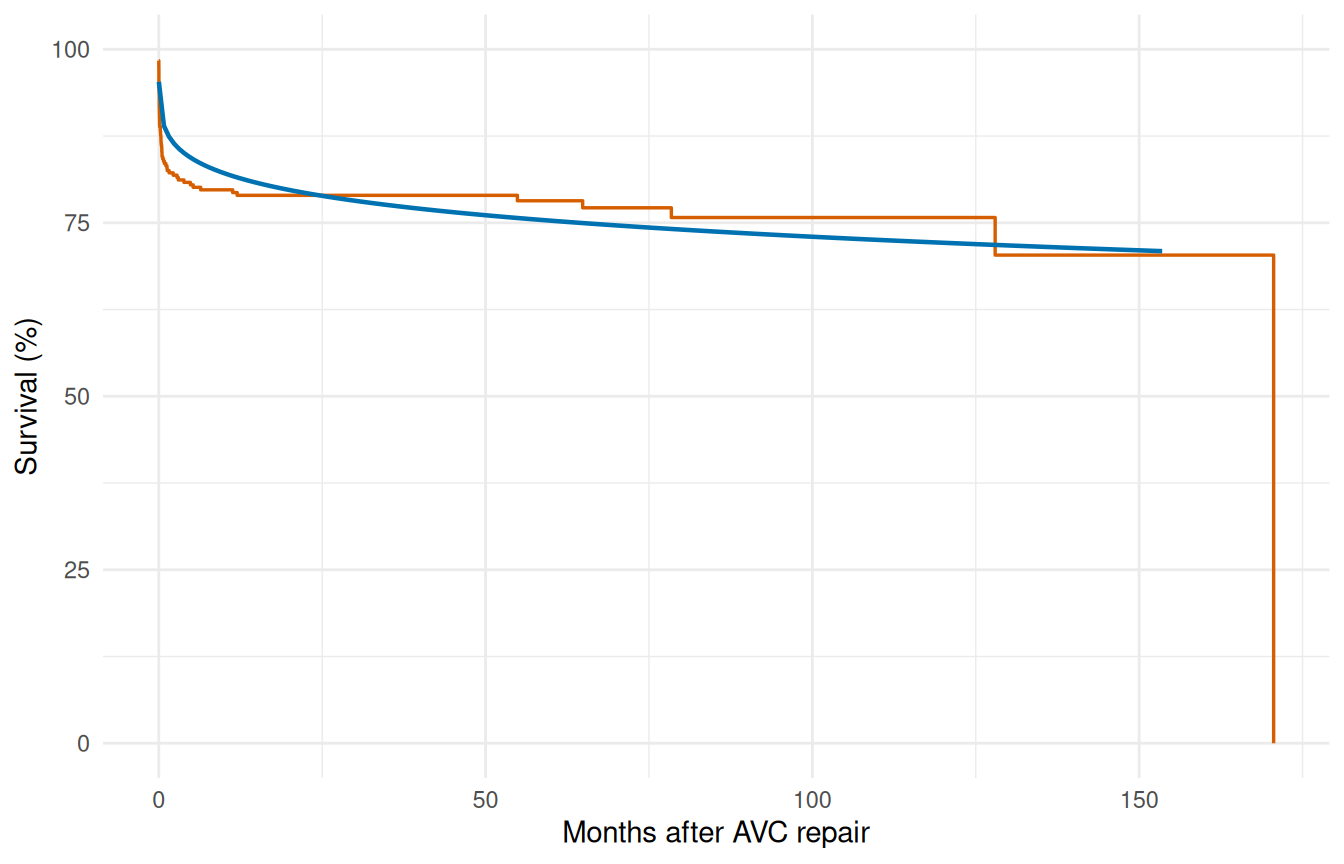
The single Weibull typically misses the sharp early drop — it compromises between the early and late time frames.
3.2 2b. Two-phase model (early CDF + constant)
The KM curve drops steeply in the first few months — operative mortality — and then settles into a roughly linear decline. The Weibull tried to capture both with one shape and ended up compromising. Splitting the hazard into two phases lets each mechanism have its own parameterization: an "cdf" early phase that saturates as operative risk resolves, plus a "constant" phase that carries the steady background attrition. Two phases is what AVC actually needs; CABG with longer follow-up would need a third late-rising phase to handle graft deterioration, but the AVC follow-up window doesn’t extend far enough to identify one.
fit_mp <- hazard(
survival::Surv(int_dead, dead) ~ 1,
data = avc,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 1, m = 1,
fixed = "shapes"),
constant = hzr_phase("constant")
),
fit = TRUE,
control = list(n_starts = 5, maxit = 1000)
)
summary(fit_mp)
#> Multiphase hazard model (2 phases)
#> observations: 305
#> predictors: 0
#> dist: multiphase
#> phase 1: early - cdf (early risk)
#> phase 2: constant - constant (flat rate)
#> engine: native-r-m2
#> converged: TRUE
#> log-lik: -228.029
#> evaluations: fn=32, gr=10
#>
#> Coefficients (internal scale):
#>
#> Phase: early (cdf)
#> estimate std_error z_stat p_value
#> log_mu -1.4132735 0.04410012 -32.04693 2.422767e-225
#> log_t_half -0.6931472 NA NA NA
#> nu 1.0000000 NA NA NA
#> m 1.0000000 NA NA NA
#>
#> Phase: constant (constant)
#> estimate std_error z_stat p_value
#> log_mu -7.609476 NA NA NANote the use of fixed = "shapes" — we fix the temporal shape parameters and only estimate the scale (log_mu) for each phase. This matches the standard HAZARD workflow: shapes are set from clinical knowledge or preliminary exploration, then scales and covariates are estimated.
surv_mp <- predict(fit_mp,
newdata = data.frame(time = t_grid),
type = "survival") * 100
ggplot() +
geom_step(data = km_df, aes(time, survival),
colour = "#D55E00", linewidth = 0.6) +
geom_line(data = data.frame(time = t_grid, survival = surv_mp),
aes(time, survival), colour = "#0072B2", linewidth = 0.8) +
labs(x = "Months after AVC repair", y = "Survival (%)") +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal()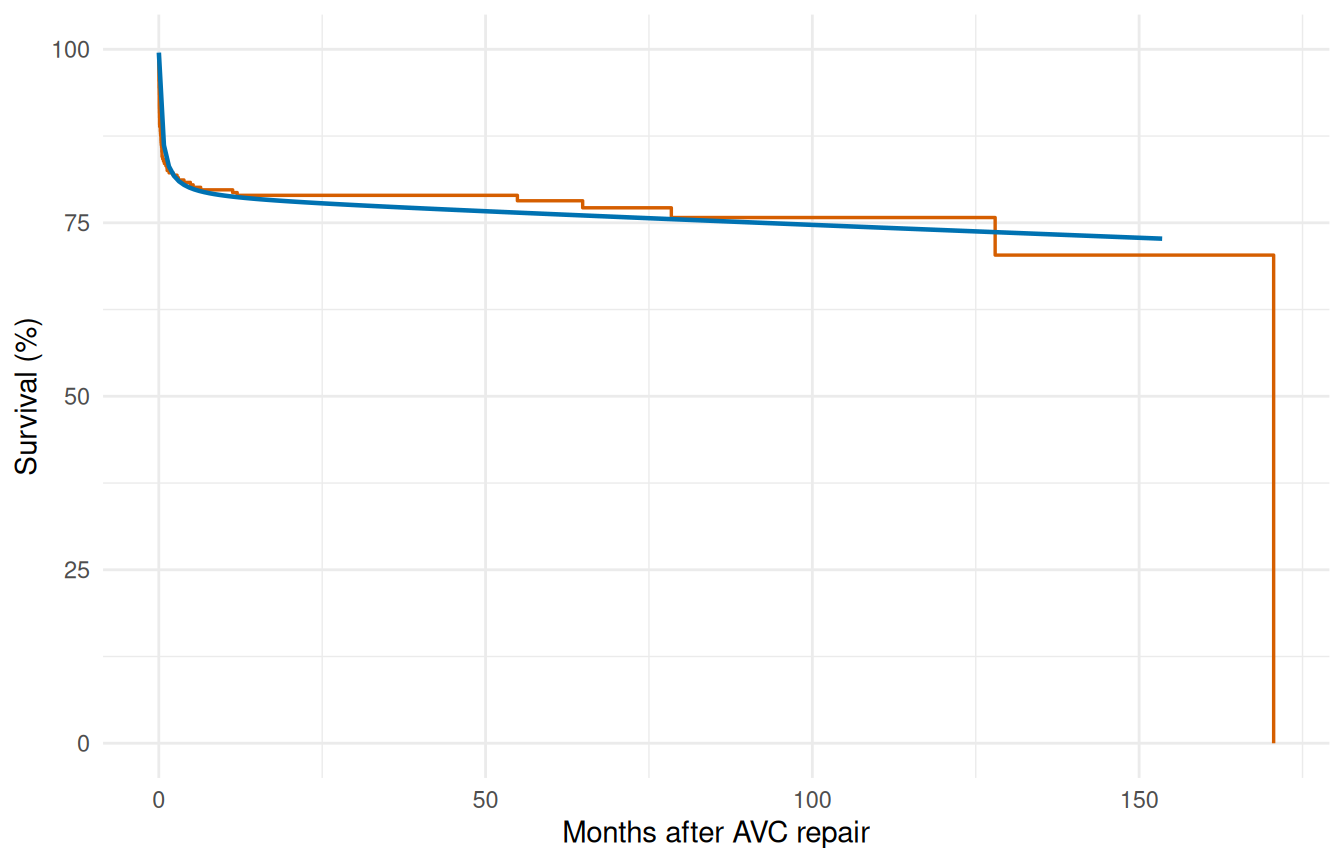
3.3 2c. Decomposed hazard
The overlay plot shows the total fit tracks KM well, but it doesn’t show whether each phase is doing the job we asked of it. Decomposing the cumulative hazard into per-phase contributions is the diagnostic that answers that question: the early phase should account for most of the steep early drop, the constant phase should carry the rest, and neither phase should be doing implausible work in the wrong time window.
decomp <- predict(fit_mp,
newdata = data.frame(time = t_grid),
type = "cumulative_hazard",
decompose = TRUE)
decomp_df <- data.frame(
time = t_grid,
total = decomp[, "total"],
early = decomp[, "early"],
const = decomp[, "constant"]
)
ggplot(decomp_df, aes(x = time)) +
geom_line(aes(y = total), linewidth = 0.9, colour = "black") +
geom_line(aes(y = early), linewidth = 0.7, colour = "#E69F00",
linetype = "dashed") +
geom_line(aes(y = const), linewidth = 0.7, colour = "#56B4E9",
linetype = "dashed") +
labs(x = "Months after AVC repair", y = "Cumulative hazard") +
theme_minimal()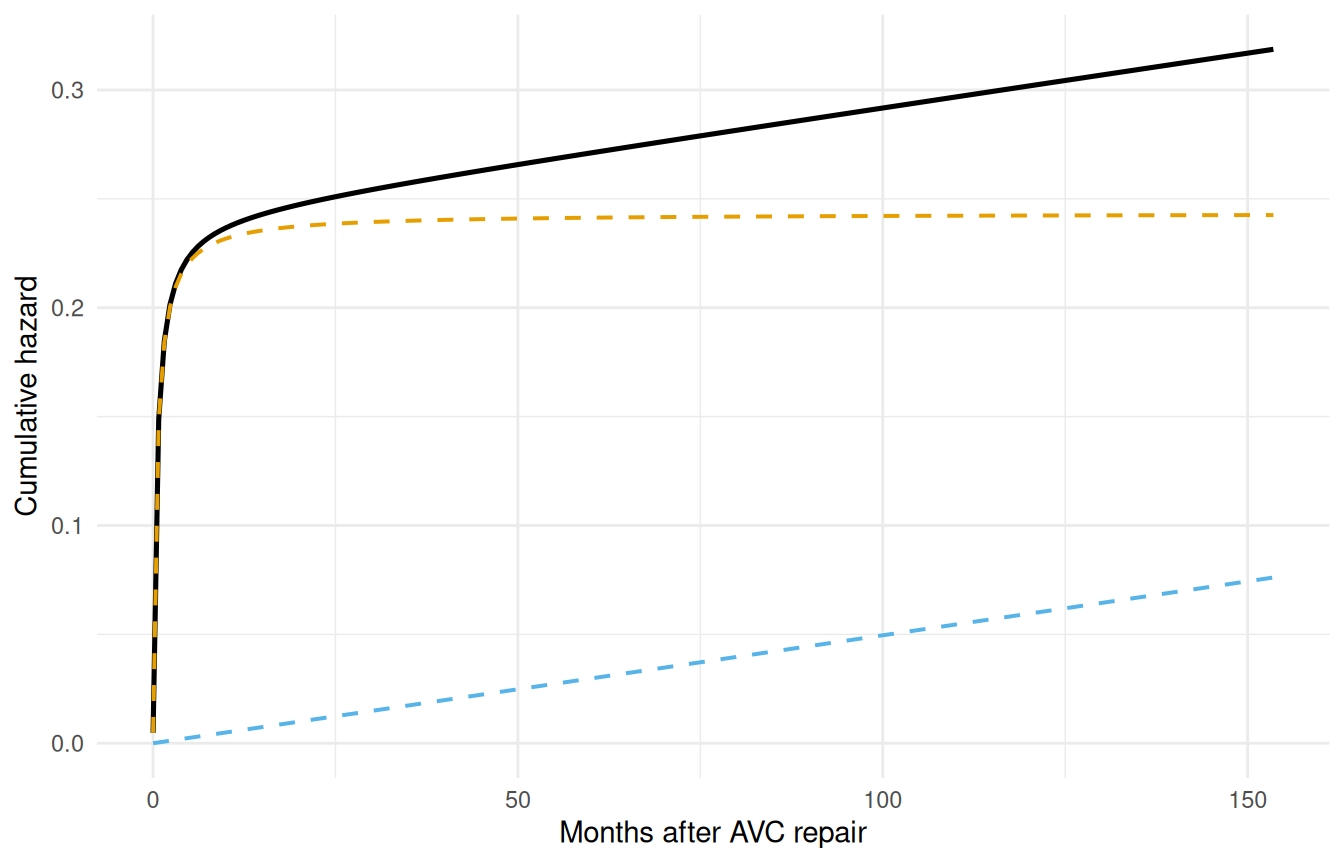
The early phase contributes most of its risk in the first few months, then levels off. The constant phase accumulates linearly over time.
4 Step 3: Variable screening
Before entering covariates into the hazard model, screen them for association with the outcome. This step corresponds to the lg.* programs in the SAS workflow.
4.1 3a. Univariable logistic screening
The cheapest screening tool we have is a univariable logistic regression of the event indicator on each covariate. It throws away the time-to-event structure but it answers a binary question quickly: does this covariate have any association with mortality at all? Covariates whose univariable p-value is huge will not suddenly become significant in the multivariable hazard model either — those can be deprioritized. Covariates with small univariable p-values deserve closer functional-form inspection before they enter the formula.
covariates <- c("age", "status", "mal", "com_iv", "orifice",
"inc_surg", "opmos")
screen_results <- data.frame(
variable = covariates,
coef = numeric(length(covariates)),
p_value = numeric(length(covariates))
)
for (i in seq_along(covariates)) {
v <- covariates[i]
fml <- as.formula(paste("dead ~", v))
lg <- glm(fml, data = avc, family = binomial)
s <- summary(lg)$coefficients
if (nrow(s) >= 2) {
screen_results$coef[i] <- s[2, 1]
screen_results$p_value[i] <- s[2, 4]
}
}
screen_results <- screen_results[order(screen_results$p_value), ]
screen_results$significant <- ifelse(screen_results$p_value < 0.05,
"*", "")
screen_results
#> variable coef p_value significant
#> 2 status 0.965232352 2.538365e-08 *
#> 4 com_iv 1.413423027 4.916970e-06 *
#> 3 mal 1.027084963 6.478986e-04 *
#> 1 age -0.004914258 1.683468e-03 *
#> 5 orifice 1.635755221 3.440489e-03 *
#> 6 inc_surg 0.293339962 1.144172e-02 *
#> 7 opmos 0.001458721 6.552347e-014.2 3b. Functional form assessment
For continuous covariates that appear significant, examine whether the relationship with outcome is monotone on the logit scale. hzr_calibrate() implements the SAS logit.sas / logitgr.sas macros: bin the continuous covariate into quantile groups, compute the observed event rate per bin, and transform it to the logit scale.
cal_age <- hzr_calibrate(x = avc$age, event = avc$dead,
groups = 10, link = "logit")
cal_age
#> Variable calibration (logit link, 10 groups)
#>
#> group n events mean min max prob link_value
#> 1 30 11 3.519 1.051 5.388 0.367 -0.547
#> 2 31 11 8.665 5.421 11.532 0.355 -0.598
#> 3 30 13 15.194 11.631 18.497 0.433 -0.268
#> 4 31 11 23.077 18.990 27.828 0.355 -0.598
#> 5 30 7 43.544 28.124 57.167 0.233 -1.190
#> 6 31 3 72.066 59.730 86.408 0.097 -2.234
#> 7 30 2 101.154 86.507 117.522 0.067 -2.639
#> 8 31 3 162.739 121.169 203.733 0.097 -2.234
#> 9 30 4 247.051 205.343 297.140 0.133 -1.872
#> 10 31 3 530.623 324.573 790.981 0.097 -2.234Plot the logit-transformed event rate against the bin centre (mean column). A straight line means the covariate can enter the model untransformed; monotone-but-curved shapes suggest a log or square-root transform; U-shapes call for a quadratic term.
ggplot(cal_age, aes(mean, link_value)) +
geom_point(size = 3, colour = "#0072B2") +
geom_line(colour = "#0072B2", alpha = 0.4) +
geom_smooth(method = "lm", formula = y ~ x, se = FALSE,
colour = "#D55E00", linetype = "dashed") +
labs(x = "Age at repair (months, decile midpoint)",
y = "logit(P(death))") +
theme_minimal()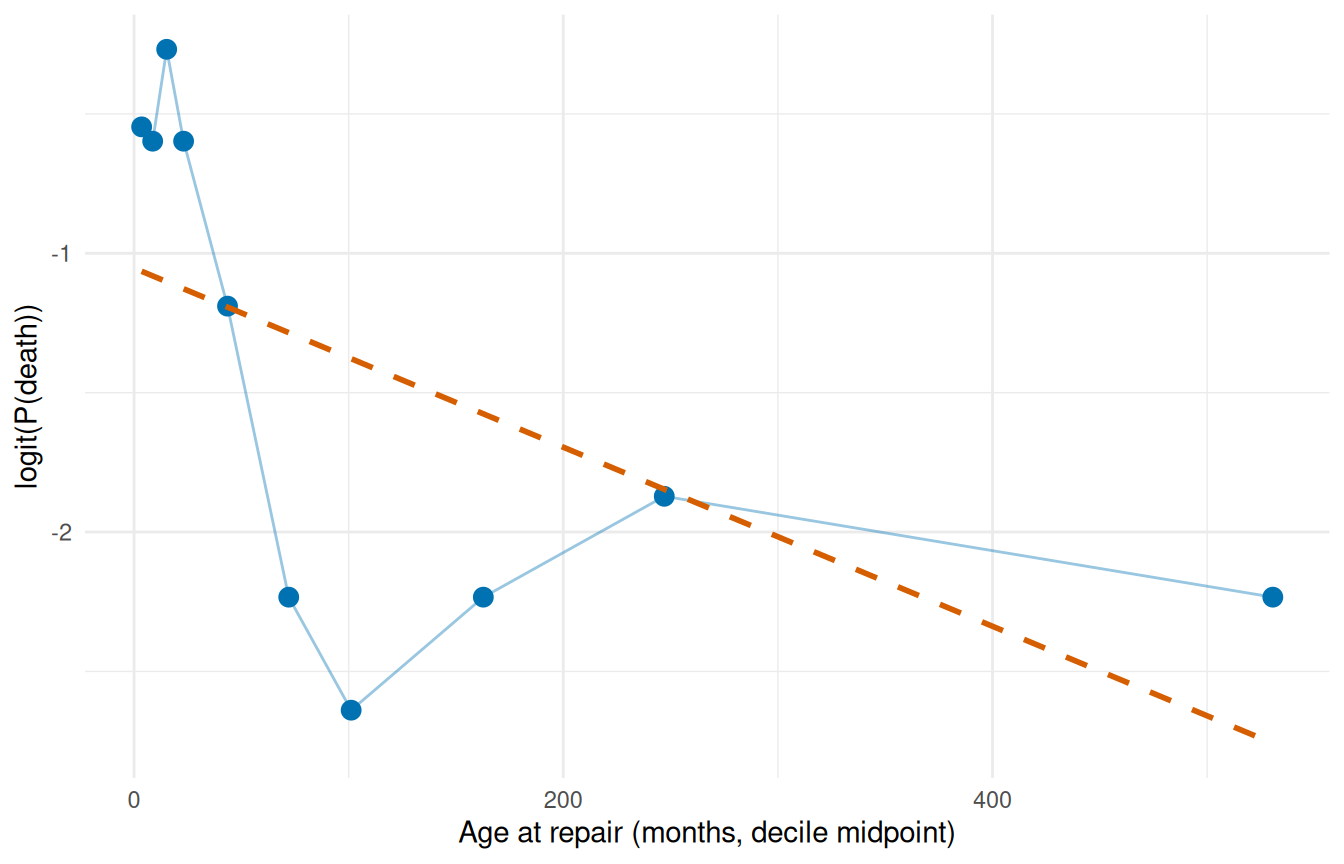
The blue points are the observed decile logits; the dashed red line is a linear reference. Deviations from linearity flag where a transform is warranted.
5 Step 4: Multivariable model
5.1 4a. Manual specification
We’ve already established the two-phase shape and screened the covariates; the multivariable model is the synthesis. Drop the covariates that passed screening onto the right-hand side of the formula and refit. The shape parameters stay fixed (we’ve earned that decision from §2); the optimizer estimates the two phase scales and one coefficient per covariate, jointly. This is the working model you’d report from this analysis.
fit_mv <- hazard(
survival::Surv(int_dead, dead) ~ age + status + mal + com_iv,
data = avc,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 1, m = 1,
fixed = "shapes"),
constant = hzr_phase("constant")
),
fit = TRUE,
control = list(n_starts = 5, maxit = 1000)
)
summary(fit_mv)
#> Warning in sqrt(d): NaNs produced
#> Multiphase hazard model (2 phases)
#> observations: 305
#> predictors: 4
#> dist: multiphase
#> phase 1: early - cdf (early risk)
#> phase 2: constant - constant (flat rate)
#> engine: native-r-m2
#> converged: TRUE
#> log-lik: -190.808
#> evaluations: fn=11, gr=1
#>
#> Coefficients (internal scale):
#>
#> Phase: early (cdf)
#> estimate std_error z_stat p_value
#> log_mu -3.605695584 NaN NA NA
#> log_t_half -0.693147181 NA NA NA
#> nu 1.000000000 NA NA NA
#> m 1.000000000 NA NA NA
#> age -0.002934522 0.00146656 -2.000956 0.045397123
#> status 0.619850638 NaN NA NA
#> mal 0.467467697 0.17964265 2.602209 0.009262544
#> com_iv 1.162696955 NaN NA NA
#>
#> Phase: constant (constant)
#> estimate std_error z_stat p_value
#> log_mu -9.6560397496 NA NA NA
#> age -0.0008940274 0.002475644 -0.3611292 0.71800288
#> status 1.0450950870 0.454449391 2.2996952 0.02146549
#> mal 0.6012599050 1.088336795 0.5524576 0.58063489
#> com_iv -1.0598349603 1.211734851 -0.8746426 0.38176838The coefficient table shows phase-specific covariate effects. A positive coefficient means higher hazard (worse prognosis). Interpretation: covariates scale the phase-specific hazard multiplicatively via exp(x * beta).
5.2 4b. Automated stepwise selection
The manual approach works when screening has already narrowed the candidate pool, but with a larger pool it helps to let the model choose. hzr_stepwise() runs forward, backward, or two-way selection against an existing hazard fit, scoring candidates with Wald p-values or delta-AIC. Defaults match SAS PROC HAZARD (SLENTRY = 0.30, SLSTAY = 0.20).
We start from a baseline with no covariates, offer the same screened candidate pool, and let the algorithm decide. For multiphase models the scope is a named list of one-sided formulas keyed by phase, so each phase can advertise its own candidate set; stepping operates per (variable, phase) pair, letting a covariate enter one phase and not another. Single-distribution models accept either a flat one-sided formula or a character vector of names.
base_mp <- hazard(
survival::Surv(int_dead, dead) ~ 1,
data = avc,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 1, m = 1,
fixed = "shapes"),
constant = hzr_phase("constant")
),
fit = TRUE,
control = list(n_starts = 3, maxit = 500)
)
fit_step <- hzr_stepwise(
base_mp,
scope = list(
early = ~ age + status + mal + com_iv,
constant = ~ age + status + mal + com_iv
),
data = avc,
direction = "both",
criterion = "wald",
trace = FALSE,
control = list(n_starts = 2, maxit = 500)
)
fit_step
#> Stepwise selection (direction = both, criterion = wald, slentry = 0.30, slstay = 0.20)
#>
#> Step 1: ENTER status into early (p = 0.000)
#> Step 2: ENTER mal into early (p = 0.000)
#> Step 3: ENTER com_iv into early (p = 0.000)
#> Step 4: ENTER age into early (p = 0.034)
#> Step 5: ENTER status into constant (p = 0.064)
#> (no further action after 5 steps)
#>
#> Final model: 5 covariates, logLik = -192.10, AIC = 396.21The $steps table records every accepted action with its Wald statistic and p-value:
fit_step$steps[, c("step_num", "action", "variable", "phase",
"p_value", "aic")]
#> step_num action variable phase p_value aic
#> 1 1 enter status early 0.000000e+00 422.9675
#> 2 2 enter mal early 1.352419e-33 416.6295
#> 3 3 enter com_iv early 1.190955e-30 399.0333
#> 4 4 enter age early 3.363966e-02 397.3463
#> 5 5 enter status constant 6.358883e-02 396.2062The final model is the fit at the top of the object, reachable through the usual hazard accessors:
logLik_manual <- fit_mv$fit$objective
logLik_step <- fit_step$fit$objective
c(manual = logLik_manual, stepwise = logLik_step,
aic_manual = 2 * length(fit_mv$fit$theta) - 2 * logLik_manual,
aic_step = 2 * length(fit_step$fit$theta) - 2 * logLik_step)
#> manual stepwise aic_manual aic_step
#> -190.8075 -192.1031 407.6150 404.2062When the screening and stepwise agree on the same covariate set the log-likelihoods match exactly; when stepwise selects a leaner model AIC drops. For an AIC-driven run use criterion = "aic"; for a forward-only sweep direction = "forward". See ?hzr_stepwise for the full argument surface including force_in, force_out, and the max_move oscillation guard.
6 Step 5: Prediction
6.1 5a. Baseline survival curve
Generate the survival curve for a reference patient (median age, NYHA class 1, no malalignment, no interventricular communication).
predict(..., se.fit = TRUE) adds a delta-method standard error and 95% confidence band around every prediction (log(-log) scale for survival so the band is guaranteed to lie in [0, 1]). This is the parametric analogue of the KM confidence limits from Step 1 — a patient-specific uncertainty estimate the nonparametric curve cannot provide.
ref_patient <- data.frame(
time = t_grid,
age = median(avc$age),
status = 1,
mal = 0,
com_iv = 0
)
surv_ref <- predict(fit_mv, newdata = ref_patient,
type = "survival", se.fit = TRUE)
ref_df <- data.frame(
time = t_grid,
survival = surv_ref$fit * 100,
lower = surv_ref$lower * 100,
upper = surv_ref$upper * 100
)
ggplot() +
geom_step(data = km_df, aes(time, survival),
colour = "#D55E00", linewidth = 0.6, alpha = 0.7) +
geom_ribbon(data = ref_df, aes(time, ymin = lower, ymax = upper),
fill = "#0072B2", alpha = 0.15) +
geom_line(data = ref_df, aes(time, survival),
colour = "#0072B2", linewidth = 0.8) +
labs(x = "Months after AVC repair", y = "Survival (%)") +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal()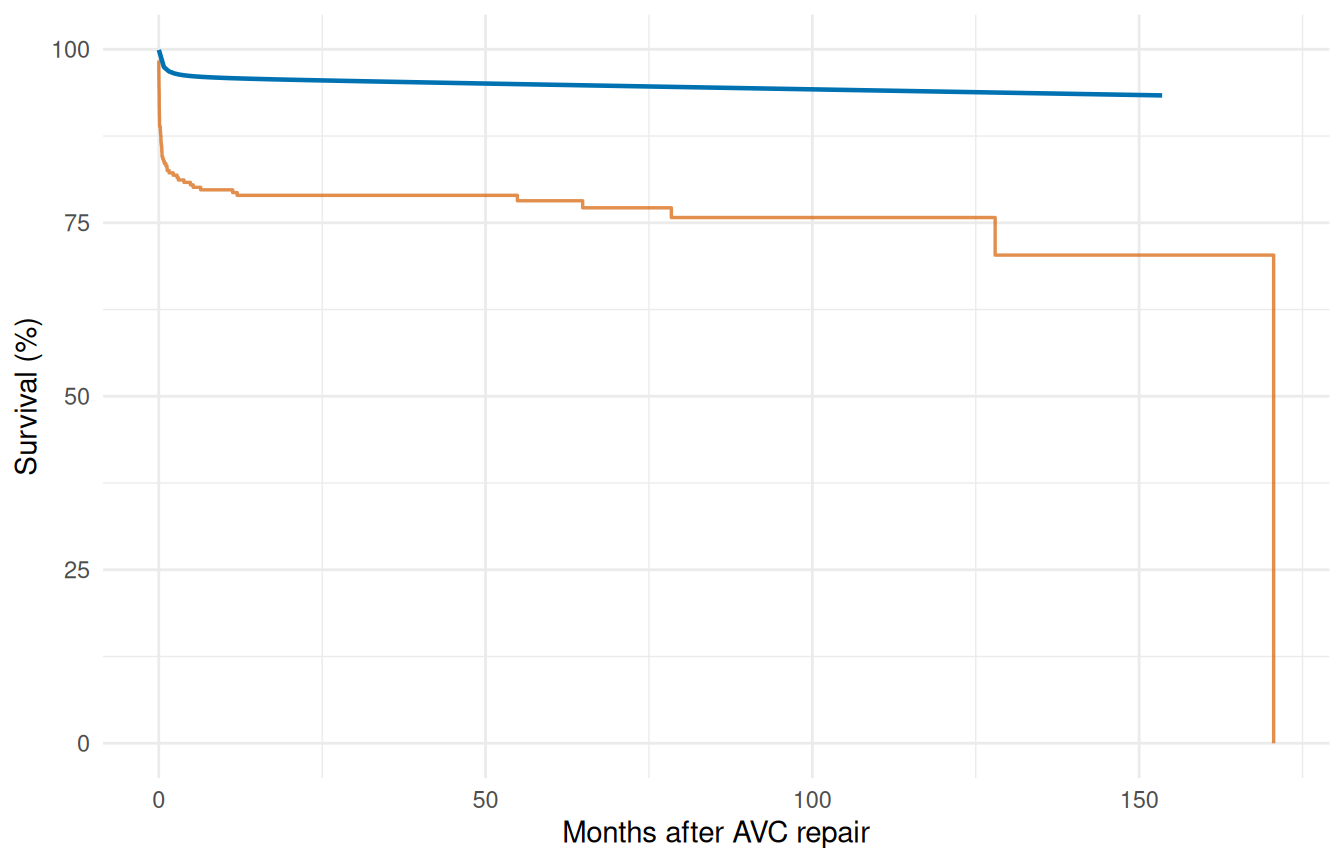
6.2 5b. Sensitivity analysis — risk factor comparison
Coefficient tables tell you about effect direction and magnitude on the log-hazard scale. They don’t directly tell a clinician what to expect at the bedside. Sensitivity analysis closes that gap by scoring two clinically meaningful profiles — a low-risk patient drawn from the favorable end of each covariate, and a high-risk patient drawn from the unfavorable end — through the fitted model and plotting the resulting survival curves with delta-method confidence bands. The vertical and horizontal gaps between the two curves are the model’s clinical-impact statement, in the units (survival probability and time) that the audience actually recognizes.
low_risk <- data.frame(
time = t_grid,
age = quantile(avc$age, 0.25),
status = 1,
mal = 0,
com_iv = 0
)
#> Warning in data.frame(time = t_grid, age = quantile(avc$age, 0.25), status = 1,
#> : row names were found from a short variable and have been discarded
high_risk <- data.frame(
time = t_grid,
age = quantile(avc$age, 0.90),
status = 4,
mal = 1,
com_iv = 1
)
#> Warning in data.frame(time = t_grid, age = quantile(avc$age, 0.9), status = 4,
#> : row names were found from a short variable and have been discarded
surv_lo <- predict(fit_mv, newdata = low_risk,
type = "survival", se.fit = TRUE)
surv_hi <- predict(fit_mv, newdata = high_risk,
type = "survival", se.fit = TRUE)
sens_df <- data.frame(
time = rep(t_grid, 2),
survival = c(surv_lo$fit, surv_hi$fit) * 100,
lower = c(surv_lo$lower, surv_hi$lower) * 100,
upper = c(surv_lo$upper, surv_hi$upper) * 100,
profile = rep(c("Low risk", "High risk"), each = length(t_grid))
)
ggplot(sens_df, aes(time, survival, colour = profile, fill = profile)) +
geom_ribbon(aes(ymin = lower, ymax = upper),
alpha = 0.15, colour = NA) +
geom_line(linewidth = 0.8) +
scale_colour_manual(values = c("Low risk" = "#0072B2",
"High risk" = "#D55E00")) +
scale_fill_manual(values = c("Low risk" = "#0072B2",
"High risk" = "#D55E00")) +
labs(x = "Months after AVC repair", y = "Survival (%)",
colour = NULL, fill = NULL) +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal() +
theme(legend.position = "bottom")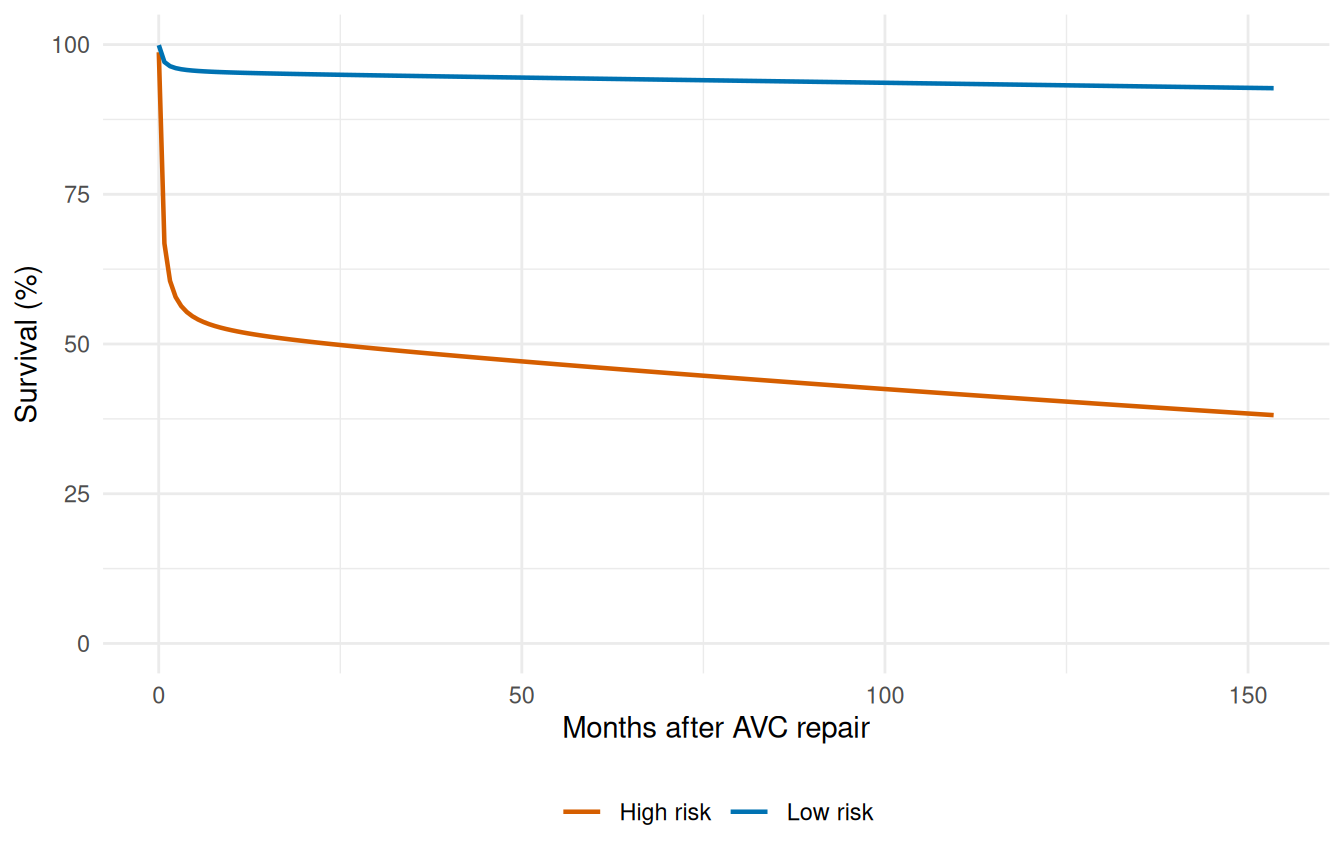
Non-overlapping confidence bands between the two profiles indicate the risk-factor combination is well-identified; overlap around the bands’ centre-of-mass flags where the model can’t distinguish the profiles with the available sample size.
7 Step 6: Validation — decile-of-risk calibration
Partition patients into deciles by predicted risk and compare observed vs. expected event rates. Good calibration means the two track each other. This implements the workflow of the SAS deciles.hazard.sas macro.
cal <- hzr_deciles(fit_mv, time = max(avc$int_dead))
print(cal)
#> Decile-of-risk calibration at time = 170.5826
#> Included 68 observations (excluded 237 censored before horizon).
#> 10 groups, 68 observed events, 29.9 expected
#>
#> group n events expected observed_rate expected_rate chi_sq p_value
#> 1 6 6 0.849 1 0.142 31.200 2.29e-08
#> 2 7 7 1.800 1 0.258 15.000 1.09e-04
#> 3 7 7 2.040 1 0.292 12.100 5.18e-04
#> 4 7 7 2.390 1 0.342 8.870 2.89e-03
#> 5 7 7 2.710 1 0.388 6.770 9.25e-03
#> 6 6 6 2.470 1 0.412 5.050 2.46e-02
#> 7 7 7 3.250 1 0.464 4.330 3.74e-02
#> 8 7 7 3.830 1 0.547 2.630 1.05e-01
#> 9 7 7 4.770 1 0.682 1.040 3.08e-01
#> 10 7 7 5.780 1 0.826 0.255 6.13e-01
#> mean_survival mean_cumhaz
#> 0.858 0.153
#> 0.742 0.298
#> 0.708 0.345
#> 0.658 0.418
#> 0.612 0.491
#> 0.588 0.530
#> 0.536 0.624
#> 0.453 0.800
#> 0.318 1.150
#> 0.174 1.820
#>
#> Overall: chi-sq = 87.2 on 9 df, p = 5.9e-15
ggplot(cal, aes(x = group)) +
geom_col(aes(y = observed_rate), fill = "#56B4E9", alpha = 0.7) +
geom_point(aes(y = expected_rate), colour = "#D55E00", size = 3) +
geom_line(aes(y = expected_rate), colour = "#D55E00", linewidth = 0.5) +
scale_x_continuous(breaks = seq_len(nrow(cal))) +
labs(x = "Risk decile (1 = lowest)", y = "Event rate",
caption = paste("Overall chi-sq =",
round(attr(cal, "overall")$chi_sq, 2),
", p =",
format.pval(attr(cal, "overall")$p_value,
digits = 3))) +
theme_minimal()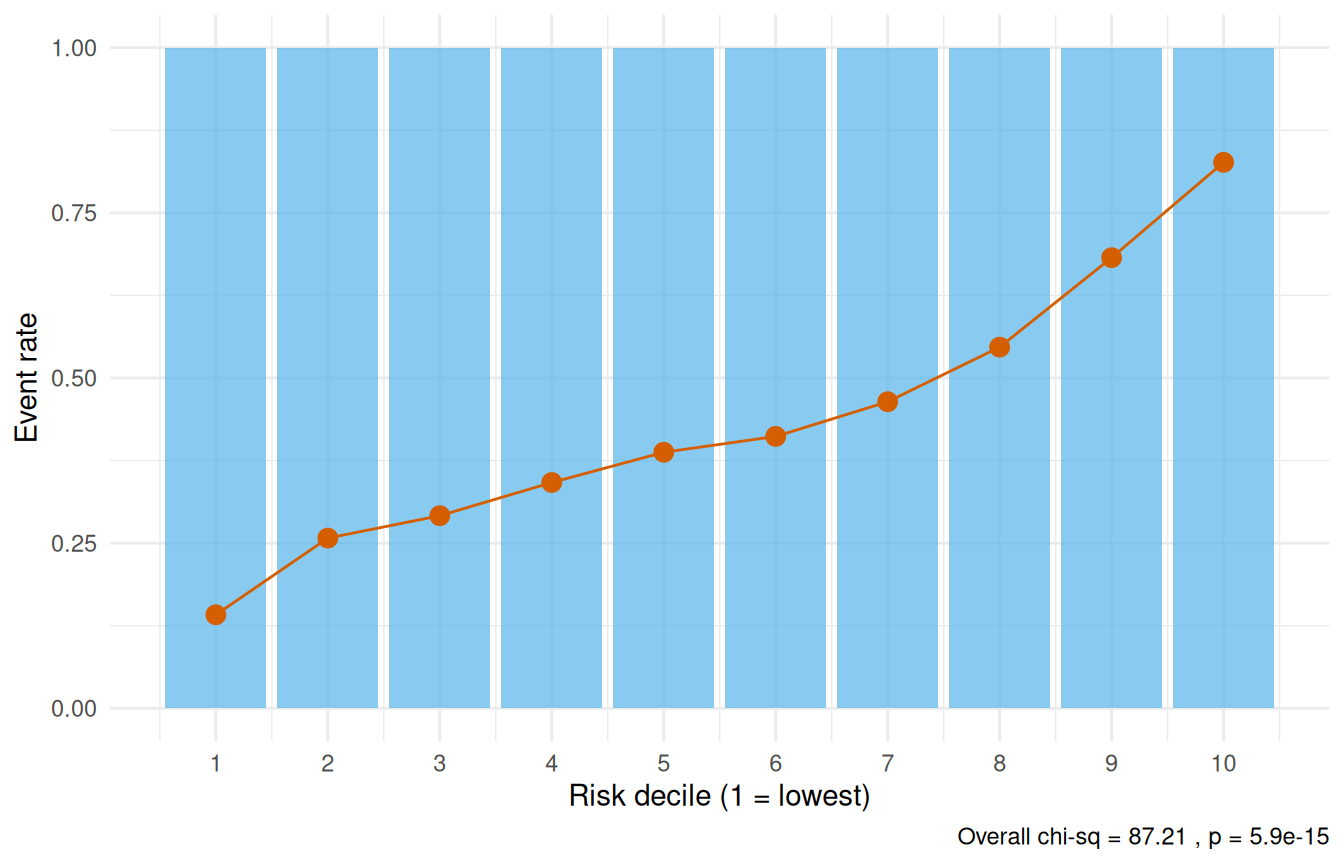
A non-significant overall chi-square statistic (p > 0.05) indicates adequate calibration — the model’s predictions are consistent with observed event rates across the risk spectrum.
7.1 Conservation of events check
The conservation-of-events principle states that a well-fitting model should predict the same total number of events as actually observed. hzr_gof() tracks this cumulatively over time — the residual (expected minus observed) should stay near zero.
gof <- hzr_gof(fit_mv)
print(gof)
#> Goodness-of-fit: observed vs. expected events
#> Distribution: multiphase | n = 305
#>
#> Total observed events: 68
#> Total expected events: 43.062
#> Final residual (E - O): -24.938
#> Conservation ratio (E/O): 0.633
#>
#> Use plot columns: time, km_surv, par_surv, cum_observed, cum_expected, residual
ggplot(gof, aes(x = time)) +
geom_step(aes(y = km_surv * 100), colour = "#D55E00", linewidth = 0.6) +
geom_line(aes(y = par_surv * 100), colour = "#0072B2", linewidth = 0.8) +
labs(x = "Months after AVC repair", y = "Survival (%)") +
coord_cartesian(ylim = c(0, 100)) +
theme_minimal()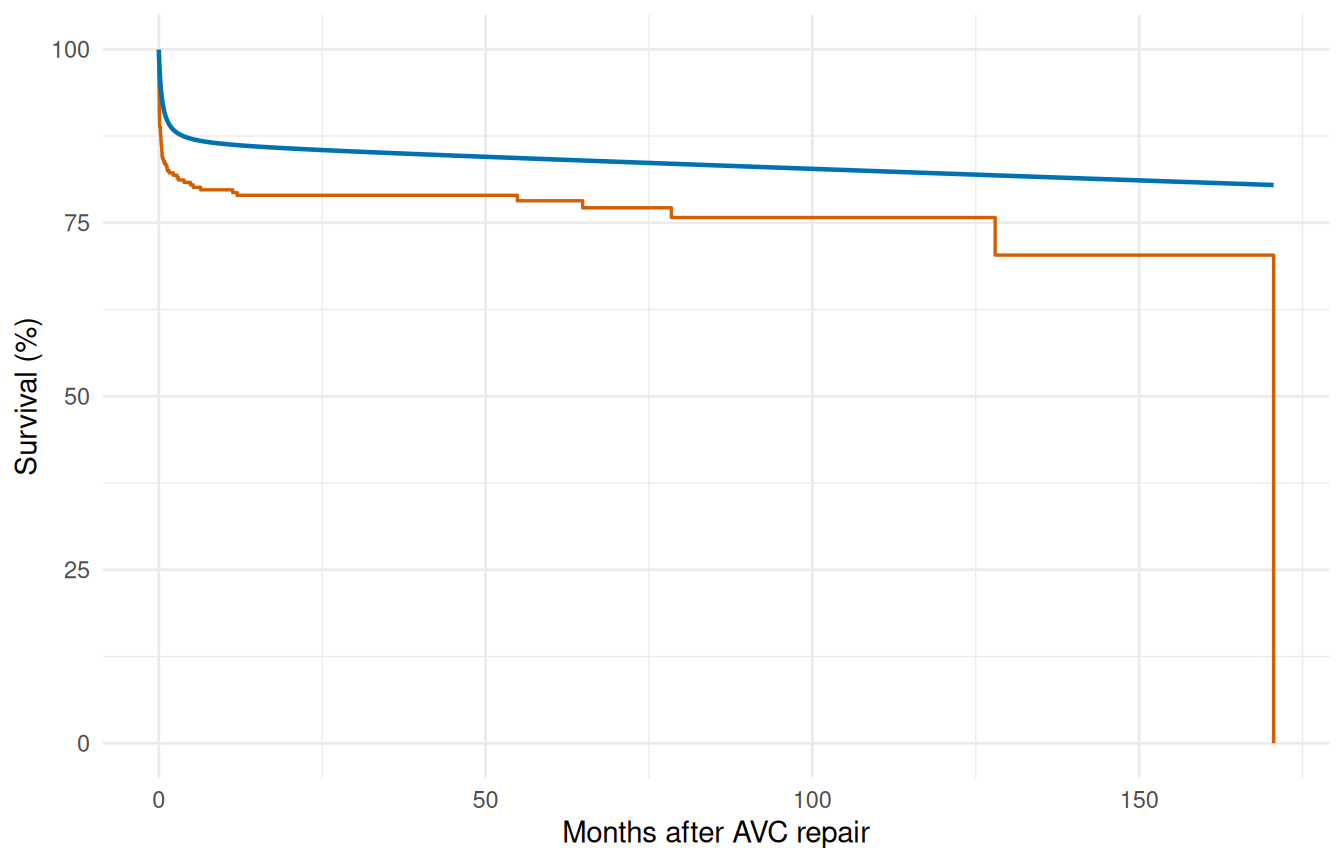
ggplot(gof, aes(x = time)) +
geom_line(aes(y = cum_observed), colour = "#D55E00", linewidth = 0.8) +
geom_line(aes(y = cum_expected), colour = "#0072B2",
linewidth = 0.8, linetype = "dashed") +
labs(x = "Months after AVC repair", y = "Cumulative events") +
theme_minimal()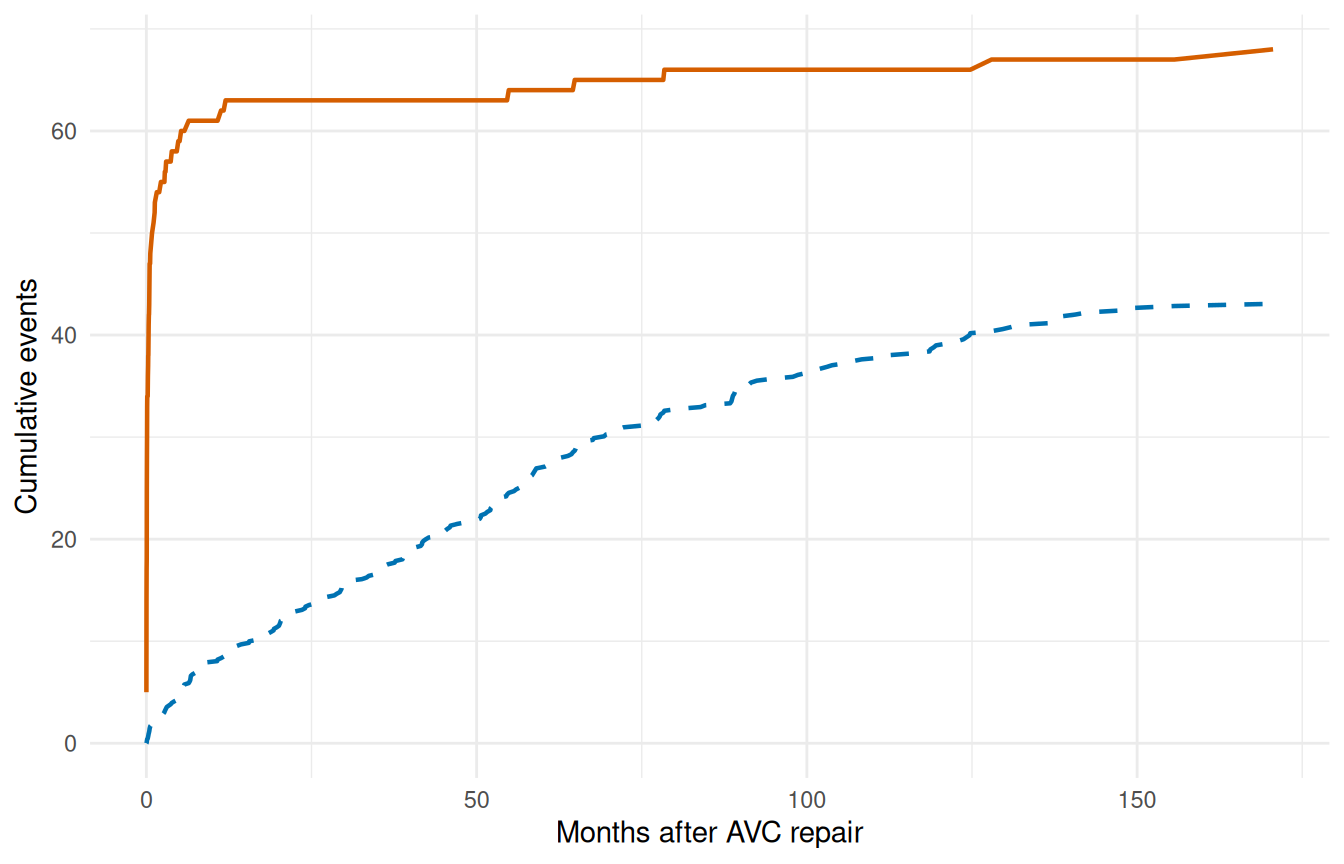
ggplot(gof, aes(x = time, y = residual)) +
geom_line(linewidth = 0.7, colour = "grey30") +
geom_hline(yintercept = 0, linetype = "dashed", colour = "grey60") +
labs(x = "Months after AVC repair",
y = "Residual (expected - observed)") +
theme_minimal()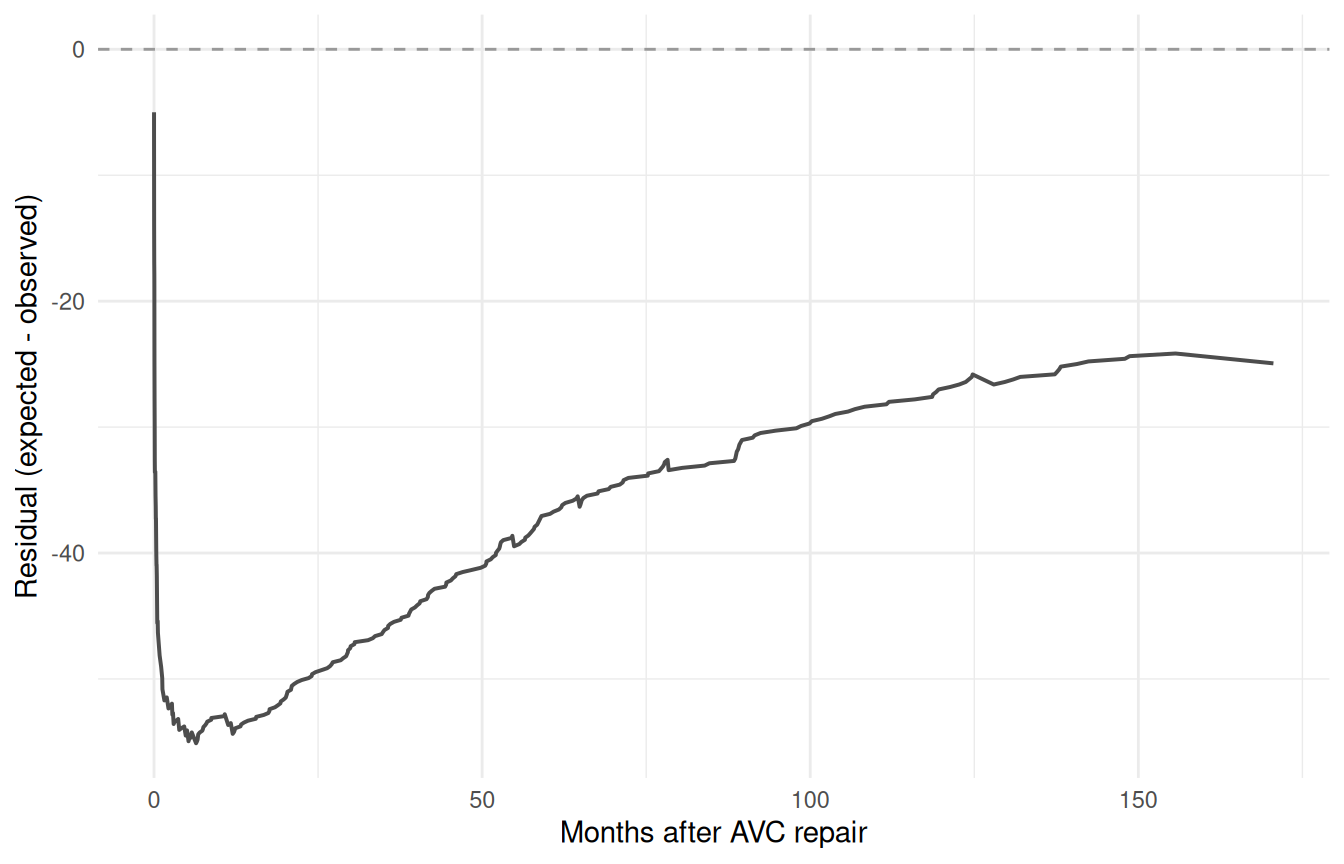
A residual that hovers near zero across the full time range indicates the model conserves events well. Persistent positive residual means the model overpredicts risk; persistent negative means it underpredicts.
8 Why not Cox regression?
For this kind of analysis the temporal parametric approach buys you several things that Cox proportional hazards does not:
| Feature | Cox (coxph) |
TemporalHazard |
|---|---|---|
| Proportional hazards required | Yes | No |
| Baseline hazard specified | No | Yes (parametric) |
| Multiple hazard phases | No | Yes (additive) |
| Patient-specific survival prediction | Approximate | Exact |
| Smooth extrapolation beyond data | No | Yes |
| Conservation of events check | No | Yes (hzr_gof()) |
| Interval censoring | Not standard | Supported |
Many clinical outcomes show non-proportional hazards: the risk factors that dominate early mortality (e.g., surgical complexity) differ from those driving late attrition (e.g., age, comorbidity). The multiphase model captures this through phase-specific covariate effects.
9 Summary
The complete analytical workflow follows a disciplined sequence:
- Kaplan-Meier baseline — establish the empirical survival pattern
- Shape fitting — match the temporal shape, simple to complex
- Variable screening — logistic screening, LOESS functional form
- Multivariable model — enter covariates with fixed shapes
- Prediction — reference curves, sensitivity analysis
- Validation — decile calibration, conservation-of-events check
This sequence ensures that the temporal shape is established before covariates are introduced, and that the final model is validated against the observed data. See vignette("getting-started") for the minimal API workflow and vignette("mf-mathematical-foundations") for the mathematical details.