Produces prediction outputs from a hazard object. Supports multiple prediction
types including linear predictor, hazard, survival probability, and cumulative hazard.
Arguments
- object
A
hazardobject.- newdata
Optional matrix or data frame of predictors. For types requiring time (e.g., "survival", "cumulative_hazard"), newdata should include a
timecolumn, or time will be taken from the fitted object's data.- type
Prediction type:
"linear_predictor": Linear predictor eta = x*beta (not available for multiphase)"hazard": Hazard scale exp(eta) (not available for multiphase)"survival": Survival probability S(t|x) = exp(-H(t|x))"cumulative_hazard": Cumulative hazard H(t|x) at event times
- decompose
Logical; if
TRUEand the model is multiphase, return a data frame with per-phase cumulative hazard contributions alongside the total. Ignored for single-distribution models. DefaultFALSE.- se.fit
Logical; if
TRUE, compute delta-method standard errors and confidence limits for each prediction. The return value becomes a data frame with columnsfit,se.fit,lower,upper. DefaultFALSE. CLs are computed on the log-hazard / log-cumhaz scale and on the log(-log(survival)) scale so lower/upper stay inside the valid range of each prediction type;linear_predictoruses symmetric natural-scale CLs. Not compatible withdecompose = TRUE.- level
Numeric confidence level in
(0, 1); default0.95. Only used whense.fit = TRUE.- ...
Unused; included for S3 compatibility.
Details
For Weibull models with survival or cumulative_hazard predictions:
Cumulative hazard: H(t|x) = (mu*t)^nu * exp(eta)
Survival: S(t|x) = exp(-H(t|x))
Time values must be positive and finite. If newdata contains a time column,
it will be used; otherwise, the time vector from the fitted object is used.
For models fit with time_windows, predictions for type = "linear_predictor"
or "hazard" also require time values (via newdata$time or fitted-time fallback)
so window-specific coefficients can be selected.
Known limitations
decompose = TRUE and se.fit = TRUE cannot be used together. The
delta-method Jacobian has not yet been extended to per-phase contributions.
To obtain confidence limits for a multiphase model, request point predictions
first (decompose = TRUE, se.fit = FALSE), then separately request CLs on
the total prediction (decompose = FALSE, se.fit = TRUE).
See also
hazard() for model fitting,
summary.hazard() for model summaries,
hzr_phase() for multiphase temporal shapes.
vignette("prediction-visualization") for detailed prediction
workflows including decomposed hazard plots and patient-specific curves.
Examples
# -- Basic predictions ------------------------------------------------
set.seed(1)
fit <- hazard(time = rexp(50, 0.3), status = rep(1L, 50),
theta = c(0.3, 1.0), dist = "weibull", fit = TRUE)
predict(fit, type = "survival")
#> [1] 0.508448836 0.315117497 0.909632142 0.913801355 0.704022509 0.034429522
#> [7] 0.297904379 0.635876619 0.407732732 0.908685617 0.245861859 0.504733067
#> [13] 0.295096862 0.003729697 0.364948062 0.373066533 0.134498594 0.565324696
#> [19] 0.772692686 0.605268524 0.070992893 0.572924045 0.803140735 0.619329758
#> [25] 0.937239852 0.968075472 0.611315229 0.007471798 0.318195648 0.389681823
#> [31] 0.232969073 0.981596565 0.781843506 0.267479469 0.868327070 0.378412625
#> [37] 0.797695039 0.524953194 0.510431884 0.845615583 0.354514703 0.376046945
#> [43] 0.276615587 0.289749395 0.626388405 0.798022001 0.276332062 0.390676459
#> [49] 0.652266897 0.113525051
predict(fit, newdata = data.frame(time = c(1, 2, 5)),
type = "cumulative_hazard")
#> [1] 0.2244716 0.5138491 1.5356737
# -- Patient-specific survival curves ---------------------------------
set.seed(1001)
n <- 180
dat <- data.frame(
time = rexp(n, rate = 0.35) + 0.05,
status = rbinom(n, size = 1, prob = 0.6),
age = rnorm(n, mean = 62, sd = 11),
nyha = sample(1:4, n, replace = TRUE),
shock = rbinom(n, size = 1, prob = 0.18)
)
fit2 <- hazard(
survival::Surv(time, status) ~ age + nyha + shock,
data = dat,
theta = c(mu = 0.25, nu = 1.10, beta1 = 0, beta2 = 0, beta3 = 0),
dist = "weibull", fit = TRUE
)
new_patients <- data.frame(
time = c(0.5, 1.5, 3.0),
age = c(50, 65, 75),
nyha = c(1, 3, 4),
shock = c(0, 0, 1)
)
# Compute predictions from the clean covariate frame before adding columns
surv <- predict(fit2, newdata = new_patients, type = "survival")
cumhaz <- predict(fit2, newdata = new_patients, type = "cumulative_hazard")
new_patients$survival <- surv
new_patients$cumulative_hazard <- cumhaz
new_patients
#> time age nyha shock survival cumulative_hazard
#> 1 0.5 50 1 0 0.9494097 0.05191482
#> 2 1.5 65 3 0 0.7743185 0.25577202
#> 3 3.0 75 4 1 0.5046535 0.68388317
# \donttest{
# -- Grouped survival curves ---------------------------------------
if (requireNamespace("ggplot2", quietly = TRUE)) {
library(ggplot2)
t_grid <- seq(0.05, max(dat$time), length.out = 80)
profiles <- data.frame(
label = c("Low risk (age 50, NYHA I)",
"High risk (age 75, NYHA IV)"),
age = c(50, 75),
nyha = c(1, 4),
shock = c(0, 1)
)
curve_list <- lapply(seq_len(nrow(profiles)), function(i) {
nd <- data.frame(
time = t_grid,
age = profiles$age[i],
nyha = profiles$nyha[i],
shock = profiles$shock[i]
)
nd$survival <- predict(fit2, newdata = nd, type = "survival") * 100
nd$profile <- profiles$label[i]
nd
})
curve_df <- do.call(rbind, curve_list)
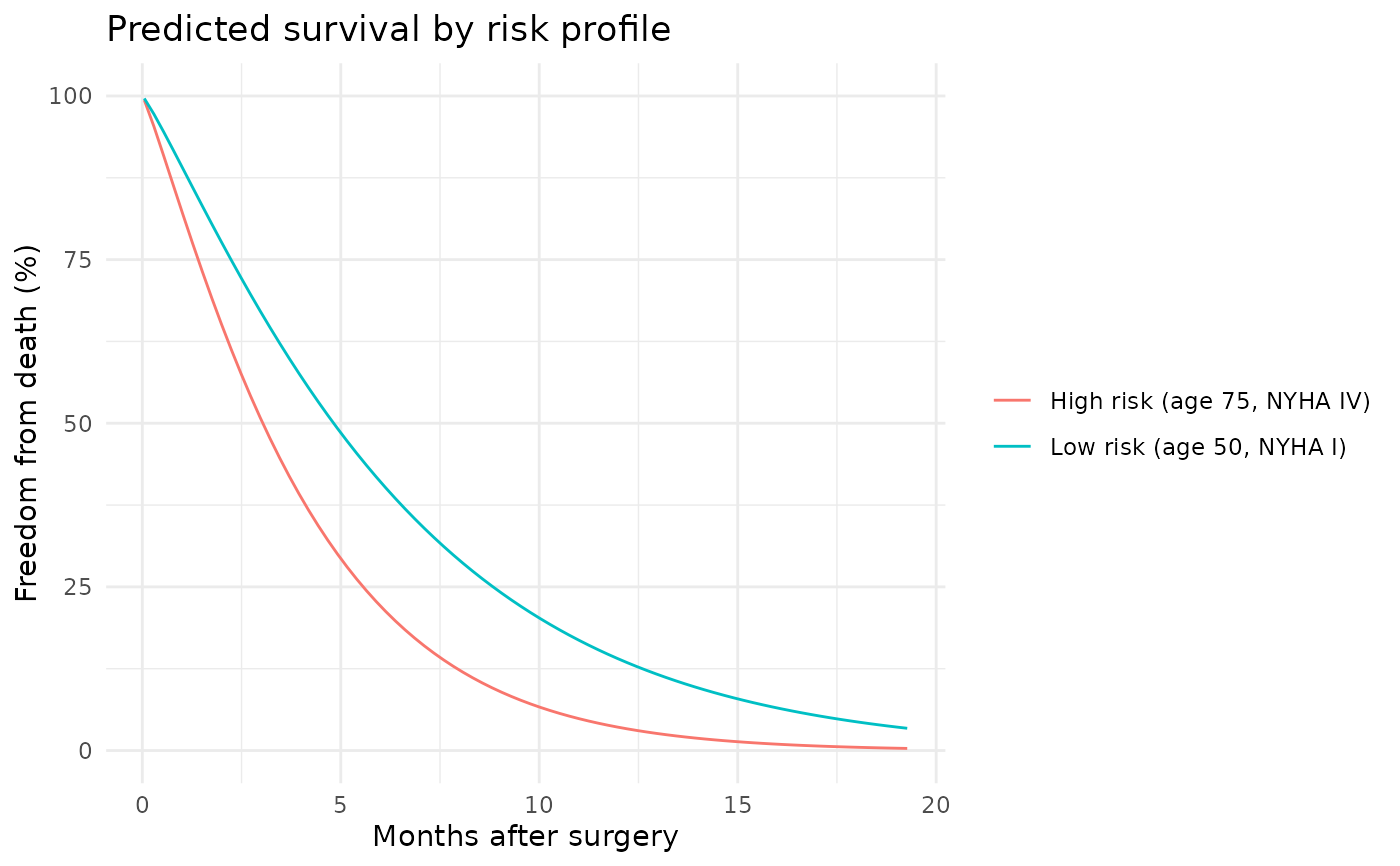
ggplot(curve_df, aes(time, survival, colour = profile)) +
geom_line() +
scale_y_continuous(limits = c(0, 100)) +
labs(x = "Months after surgery",
y = "Freedom from death (%)",
title = "Predicted survival by risk profile",
colour = NULL) +
theme_minimal()
}
 # }
# \donttest{
# -- Multiphase predictions with decomposition --------------------
set.seed(42)
n <- 200
dat <- data.frame(
time = rexp(n, rate = 0.25) + 0.01,
status = rbinom(n, size = 1, prob = 0.65)
)
fit_mp <- hazard(
survival::Surv(time, status) ~ 1,
data = dat,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 2, m = 0,
fixed = "shapes"),
late = hzr_phase("cdf", t_half = 5, nu = 1, m = 0,
fixed = "shapes")
),
fit = TRUE,
control = list(n_starts = 5, maxit = 1000)
)
t_grid <- seq(0.01, max(dat$time) * 0.9, length.out = 100)
nd <- data.frame(time = t_grid)
# Overall survival
predict(fit_mp, newdata = nd, type = "survival")
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.9991035 0.9506793 0.9308162 0.8914472 0.8319967 0.7661083
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.7030012 0.6464729 0.5973334 0.5551219 0.5189604 0.4879204
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.4611599 0.4379618 0.4177323 0.3999852 0.3843243 0.3704264
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.3580275 0.3469102 0.3368953 0.3278339 0.3196015 0.3120937
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.3052222 0.2989121 0.2930993 0.2877292 0.2827542 0.2781335
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2738314 0.2698169 0.2660626 0.2625446 0.2592417 0.2561351
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2532082 0.2504459 0.2478352 0.2453639 0.2430214 0.2407981
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2386851 0.2366746 0.2347594 0.2329329 0.2311892 0.2295228
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2279288 0.2264026 0.2249400 0.2235372 0.2221905 0.2208967
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2196529 0.2184561 0.2173037 0.2161934 0.2151230 0.2140902
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2130932 0.2121302 0.2111994 0.2102993 0.2094284 0.2085853
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2077688 0.2069774 0.2062103 0.2054661 0.2047440 0.2040429
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2033620 0.2027005 0.2020574 0.2014320 0.2008237 0.2002317
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1996554 0.1990942 0.1985474 0.1980146 0.1974952 0.1969888
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1964947 0.1960127 0.1955422 0.1950828 0.1946343 0.1941961
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1937679 0.1933494 0.1929403 0.1925402 0.1921489 0.1917661
#> early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1913914 0.1910247 0.1906657 0.1903142
# Per-phase decomposed cumulative hazard
decomp <- predict(fit_mp, newdata = nd,
type = "cumulative_hazard", decompose = TRUE)
head(decomp)
#> time total early late
#> 1 0.0100000 0.0008968715 0.0008968715 5.301145e-151
#> 2 0.3177112 0.0505785158 0.0505467552 3.176063e-05
#> 3 0.6254224 0.0716934643 0.0648896246 6.803840e-03
#> 4 0.9331336 0.1149091204 0.0726070448 4.230208e-02
#> 5 1.2408448 0.1839268533 0.0776684396 1.062584e-01
#> 6 1.5485560 0.2664317832 0.0813355656 1.850962e-01
# }
# }
# \donttest{
# -- Multiphase predictions with decomposition --------------------
set.seed(42)
n <- 200
dat <- data.frame(
time = rexp(n, rate = 0.25) + 0.01,
status = rbinom(n, size = 1, prob = 0.65)
)
fit_mp <- hazard(
survival::Surv(time, status) ~ 1,
data = dat,
dist = "multiphase",
phases = list(
early = hzr_phase("cdf", t_half = 0.5, nu = 2, m = 0,
fixed = "shapes"),
late = hzr_phase("cdf", t_half = 5, nu = 1, m = 0,
fixed = "shapes")
),
fit = TRUE,
control = list(n_starts = 5, maxit = 1000)
)
t_grid <- seq(0.01, max(dat$time) * 0.9, length.out = 100)
nd <- data.frame(time = t_grid)
# Overall survival
predict(fit_mp, newdata = nd, type = "survival")
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.9991035 0.9506793 0.9308162 0.8914472 0.8319967 0.7661083
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.7030012 0.6464729 0.5973334 0.5551219 0.5189604 0.4879204
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.4611599 0.4379618 0.4177323 0.3999852 0.3843243 0.3704264
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.3580275 0.3469102 0.3368953 0.3278339 0.3196015 0.3120937
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.3052222 0.2989121 0.2930993 0.2877292 0.2827542 0.2781335
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2738314 0.2698169 0.2660626 0.2625446 0.2592417 0.2561351
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2532082 0.2504459 0.2478352 0.2453639 0.2430214 0.2407981
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2386851 0.2366746 0.2347594 0.2329329 0.2311892 0.2295228
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2279288 0.2264026 0.2249400 0.2235372 0.2221905 0.2208967
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2196529 0.2184561 0.2173037 0.2161934 0.2151230 0.2140902
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2130932 0.2121302 0.2111994 0.2102993 0.2094284 0.2085853
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2077688 0.2069774 0.2062103 0.2054661 0.2047440 0.2040429
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.2033620 0.2027005 0.2020574 0.2014320 0.2008237 0.2002317
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1996554 0.1990942 0.1985474 0.1980146 0.1974952 0.1969888
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1964947 0.1960127 0.1955422 0.1950828 0.1946343 0.1941961
#> early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1937679 0.1933494 0.1929403 0.1925402 0.1921489 0.1917661
#> early.log_mu early.log_mu early.log_mu early.log_mu
#> 0.1913914 0.1910247 0.1906657 0.1903142
# Per-phase decomposed cumulative hazard
decomp <- predict(fit_mp, newdata = nd,
type = "cumulative_hazard", decompose = TRUE)
head(decomp)
#> time total early late
#> 1 0.0100000 0.0008968715 0.0008968715 5.301145e-151
#> 2 0.3177112 0.0505785158 0.0505467552 3.176063e-05
#> 3 0.6254224 0.0716934643 0.0648896246 6.803840e-03
#> 4 0.9331336 0.1149091204 0.0726070448 4.230208e-02
#> 5 1.2408448 0.1839268533 0.0776684396 1.062584e-01
#> 6 1.5485560 0.2664317832 0.0813355656 1.850962e-01
# }